When I see limitations in published network papers that I find sufficiently relevant to address, I discuss the papers here, and try to provide ways forward. For instance, Madhoo & Levine (2016) drew inferences from visually inspecting their networks, which inspired me to write a tutorial blog post on the topic. And when the paper by Afzali et al. (2016) could have benefitted from additional stability analyses, and I wrote a tutorial blog post on network stability.
Today’s blog tackles the replicability of network models, and I will provide my personal take on the topic here. The blog does not reflect the view of my colleagues or co-authors1, and is a very personal tale … full of woe and wonder.
The Journal of Abnormal Psychology just decided to go ahead with publishing a paper although there is at least one known serious error, and a number of major problems. I read this paper, entitled “Evidence that psychopathology networks do not replicate”, the first time about half a year ago (the title changed since), when we were invited by Abnormal to write a commentary. In the paper, Forbes, Wright, Markon and Krueger, from here on FWMK, fit 4 network models to two large community datasets of 18 depression and anxiety symptoms, and investigated whether the network models replicate across the 2 datasets.
FWMK conclude:
“Popular network analysis methods produce unreliable results”, “Psychopathology networks have limited replicability”, “poor utility”, and, later, “current psychopathology network methodologies are plagued with substantial flaws”.
That is devastating, and is worrisome for anybody who has used network models before. So we — Denny Borsboom, Sacha Epskamp, Lourens Waldorp, Sacha Epskamp, Claudia van Borkulo, Han van der Maas, Angelique Cramer, and me — sat down and took a closer look at the paper, and found the following problems. And just to highlight that again, I’m describing the version of the paper we received after it was accepted for publication.
- For the Directed Acyclic Graphs (DAGs), the authors had used the correlation matrix instead of the raw data as input for the R-package bnlearn, leading to nonsensical results.
- For the relative importance networks, FWMK had deleted the strongest edges from the estimated networks without mentioning this in the paper itself. This procedure makes no sense, nobody has done that before, and it is akin to deleting the strongest regression coefficients from a regression, or the strongest factor loadings from a factor model, and not reporting that you do so in the paper.
- The authors fit these relative importance networks that are based linear regressions to binary symptom data, which violates basic distributional assumptions; again, nobody in the literature has done that before.
- For the association networks (simple visualizations of the correlation matrices), they had based their analysis on non positive definitive correlation matrices without acknowledging it in the paper. The correlation matrix was non positive definitive because the authors imputed a ton of missing data based on skip questions with 0s, dramatically distorting the correlation matrix. The distortion is so severe that the average correlation among depression items was 0.33 before imputation (after listwise deletion), and 0.95 after imputation.
- Only 25% of the four models — the Ising Models — were estimated correctly.
This is a remarkable collection of issues for a paper that draws the strong conclusions about methodology I quoted above, and took us a considerable amount of time to identify all by digging through the code of FWMK. We informed the editor that the paper contains a number of serious flaws, and were surprised that Abnormal decided to go along with the publication of the paper. We followed the editor’s invitation to write a commentary, in which we re-analyzed the data and fixed the mistakes we identified; all code and results are available online.
A number of curious things happened next. First, because the editor gave us only a few weeks to reply, and because we wanted to re-analyze the data to make sense of the implausible network structures in the manuscript, we asked the editor to confirm in writing before we started working on the commentary that this is the final accepted version of the manuscript; the editor confirmed it is.
Second, we then found out that we have to apply for one of the two datasets, and pay for it, because the replicability paper by FWMK with very strong claims about a whole family of psychometric models was actually not reproducible2.
Third, while working on our re-analysis, FWMK changed the final accepted version of the manuscript (that we were guaranteed by the editor to be final) not once, but twice3. In total, FWMK fixed the DAG errors, rewrote parts of the paper, changed the title (“Evidence that psychopathology networks have limited replicability”) and the results, but left the discussion and conclusions untouched. The paper was not peer-reviewed after the changes4, and the incorrect estimation of the the relative importance networks persisted5, so did association networks based on non positive definitive correlation matrices, implausible correlations among depression symptoms of 0.95 due to zero-imputation, and the application of linear regression to binary data. And, of course, the devastating conclusions about network methodology in general.
Let’s ignore the fact that the editor refused to give us even one more week to write the commentary, despite the authors changing the paper twice while we were writing up the commentary and re-analyzing all data. And let’s ignore the fact that the editor and the reviewers insisted we cannot call any aspect of the manuscript an “error” or “wrong”, but needed to use words such as “statistical inaccuracy” — while they were happy to have FWMK draw extremely strong conclusions about methodology based on (mis)applying models to two datasets. And let’s also ignore for now that the editor asked specifically us to declare conflicts of interests because we “teach network models” — despite the fact that FWMK also teach (e.g. factor models), despite the fact that Steinley et al. (also invited to comment on FWMK’s paper) also teach network analysis, and despite the fact that it is fairly uncommon to read: “We report severe conflicts of interests regarding the t-tests used in this manuscript … because we teach them to students.”6
In any case, ignoring the very weird review and publication process, the main point here is that the Journal of Abnormal Psychology and FWMK decided to go ahead with publishing a paper that contained significant errors that the editor and the authors were aware of.
In the final paper that was published yesterday …
- … 25% of the models (relative importance networks) are wrong
- … 25% of the models (the association networks) are based on non positive definitive correlation matrices with highly implausible values
- … 25% of the models (DAGs) were re-estimated after final acceptance of the paper, and the respective sections rewritten, without peer-review
- … and 25% of the models (Ising Models) do not contain the only validated statistical test for assessing replicability (the Network Comparison Test); using this test in their data leads to the opposite conclusion FWMK draw.
These facts are not available to readers of the paper.
Non sequitur
I haven’t addressed the biggest problem yet. Assume we don’t understand regressions very well: it is a new methodology. You fit a regression of smoking on mortality to a large community dataset 1, and then you fit the same regression to a large community dataset 2. The coefficients are very similar. You write this up and call your paper “Regression methodology replicates well”.
Now imagine the opposite: the coefficients are very different across the two datasets, and you call your paper “Regression methodology does not replicate”.
Both conclusions are equally absurd. Why? Because you cannot vet the methodology of regression analysis by applying it to two different datasets. After all, the results could be different because the datasets differ, and you might find different results if you look into different data. You need simulation studies for that7.
FWMK published pretty much that paper, except that they fit network models, and not regressions, to two datasets, and then draw conclusions about network methodology. Actually, they kind of did publish that paper, because network models are a bunch of regressions, with a bit of regularization on top.
Irrespective of the results FWMK obtained, conclusions about methodology do not follow from fitting models to two datasets — because you do not know the true model in these datasets, which could differ. To vet methodology, you want to simulate data from a given true model and see if you can estimate it back reliably with your statistical procedure.
This point — vetting methodology requires more than fitting a method to two datasets — is one of the main points we made in our invited rejoinder. Unfortunately, the authors did not address this point in their rebuttal. To see how weird the claim is that network methodology is flawed because it does not replicate across 2 datasets, let’s exchange “network model” with “factor model”, and fit 12 different factor models for the MADRS depression scale established in the prior literature to a new dataset (Quilty et al., 2013). We find that only 1 of these 12 models provides acceptable fit. If FWMK read that paper, would they conclude that “factor models do not replicate” and are “plagued by substantial flaws”? Of course not. They would, like me, conclude that these datasets really seem to differ in the correlation matrices and factor structures. This is not a shortcoming of factor models, but due to differences in data.
Do networks replicate or not?
The conclusions about network models in general the authors draw from fitting network models to two datasets do not follow. But we can ask the question instead: how well do network models replicate across these two specific datasets FWMK used, similar to asking: how well do factor models replicate across these two specific datasets? This is an interesting empirical question, which fits the outlet of the paper, an applied clinical journal.
Network models — just like factor models — produce parameters, and the question of replicability is how similar these parameters are across the models fit to two datasets. An Ising Model, for instance, has (k * (k-1)) / 2 edge parameters (where k is the number of items)8, so in the case of the FWMK data with 18 items, 153 parameters.
There are many ways to compare these 153 parameters across two models. The quickest (and dirtiest) way is to correlate these parameters. FWMK fit three network models (the Association Network is simply a visualization of the correlation matrix, which we won’t count as a ‘model’ here): Ising Models, Relative Importance Networks, and DAGs. The correlations of parameters are 0.95 for the Ising Model, and 0.98 for the relative importance network9. These correlations are not provided in the original paper, the authors instead report the % of edges of networks models in dataset 1 that are also identified in the models fitted to dataset 2: 86.3% for the Ising Model, 98.3% for the relative importance network10, and 79.4% for the DAGs.
Even if the authors hadn’t made mistakes, and even if we take their results at face value — I fail to understand how FWMK went from their own results to calling their paper “Evidence that psychopathology networks have limited replicability”. In our re-analysis of the data, we used some additional metrics to assess replicability in addition to correlations of parameters and % of edges that replicate, all of which are reported in the main table of our rejoinder.
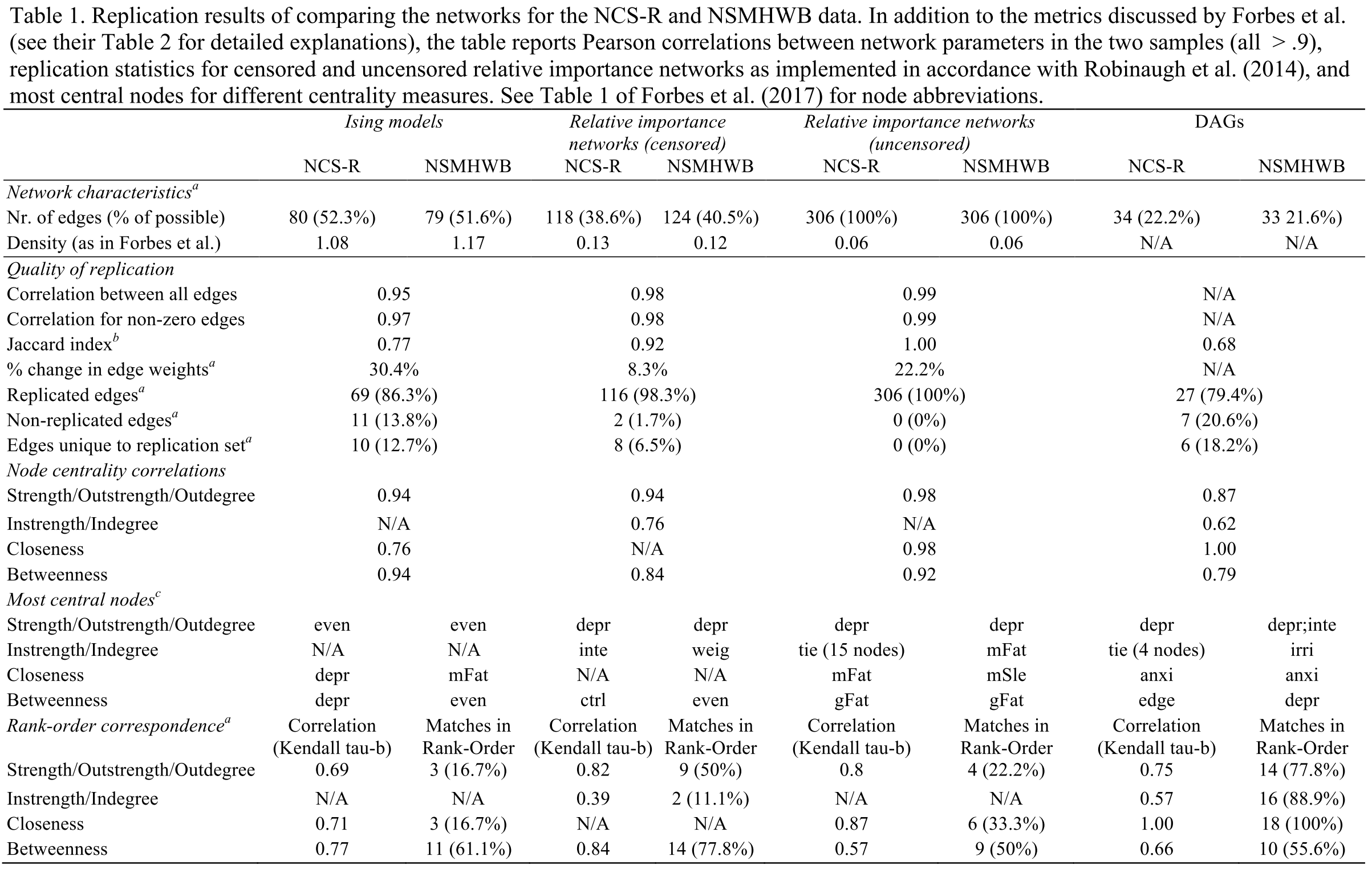
Of note, we also used the Network Comparison Test, a validated statistical test to compare Ising Models across datasets; you can think about this as being similar to measurement invariance tests in the factor modeling literature11. The result of the test was that no significant difference between the two Ising Models could be detected, which was expected given the very high correlation among parameters, and the very high replicability of individual edges.
Skip imputation
Now, I couldn’t stress enough — and we do so in the commentary as well — that the high replicability of network models in these two datasets does not make network models great, or replicable. In fact, it says very little about network models in general — that’s what simulation studies are for — and we conclude in our commentary that the stunning similarity of the network models comes from the skip imputation the authors performed. In my own work on network replicability (4 clinical datasets of patients receiving treatment for PTSD, no skip questions), the similarity of network structures is somewhat lower than in FWMK … but more of that later.
The authors used two datasets that had skip questions for anxiety and depression symptoms. For instance, symptoms 3 to 9 for Major Depression were not coded if people did not have at least symptoms 1 or 2. These missing values are commonly replaced with 0s, which is what FWMK also did. In both datasets. You see where this is going: you induce the same spurious correlations in both datasets and then assess the replicability of statistical models that rely on the (partial) correlations among items. This makes investigating replicability very difficult, because you cannot distinguish the signal in the data from the spurious associations induced by replacing skip-out items by zeros. In the data of FWMK, listwise deletion leads to correlation coefficients of 0.33 in the data12, while zero-imputation leads to a non positive definite correlation matrix, the next positive definite of which features average correlations of 0.95. This is not a plausible correlation matrix.
Now, is it ok to impute skip missing data with 0s? I cannot answer that question here in general. Is it commonplace? Absolutely. Does it make sense to induce spurious correlations in two datasets at the same time when you want to compare how well a statistical model based on item covariances generalizes from one dataset to the other? It is a very big problem. FWMK not only ignored the topic completely in their original paper, but conclude in their rebuttal that “zero-imputation is thus a potential limitation of extant network approaches”. But obviously, factor or IRT models, and even regressions, would have exactly the same issues: if you replace missings on items 3 to 9 by 0s in case people do not have item 1 or 2, you will create spurious dependencies among items 3 to 9 (because they often get 0s together), and you will also create spurious dependencies between items 3 to 9 and items 1 and 2 (because 3 – 9 depend on the presence of 1 and 2). Concluding that “zero-imputation is thus a potential limitation of regression analysis” would be equally silly as the conclusion FWMK draw. The authors’ rebuttal that other network papers in the past have based the estimation of network models on data after zero imputation does not change the fact that they ignored an issue that was discussed already in the very first empirical network paper by the Amsterdam Psychosystems group13 and several other papers14, that the strategy altered the correlation among items in their datasets dramatically, and that the strategy introduced the same spurious relationships among the two datasets they wanted to compare. In addition, other researchers who used the approach, like Borsboom & Cramer15, clearly stated: “The emphasis on free availability of data and replicability of the reported analyses occasionally means that the analyses may not be fully appropriate for the data (e.g., when computing partial correlations on dichotomous variables); in these cases, which will be indicated to the reader, the empirical results have the main purpose of illustration rather than interpretation in meaningful substantive terms.” This differs from the devastating conclusions FWMK draw about a whole family of statistical models.
Visualization
Another point I find important to highlight is that FWMK use a different layout for the networks to show how different they are. To show you why this is a problem, let me give you an example: are the two network models below — that I just made up, they don’t have anything to do with the results of FWMK — the same or not?
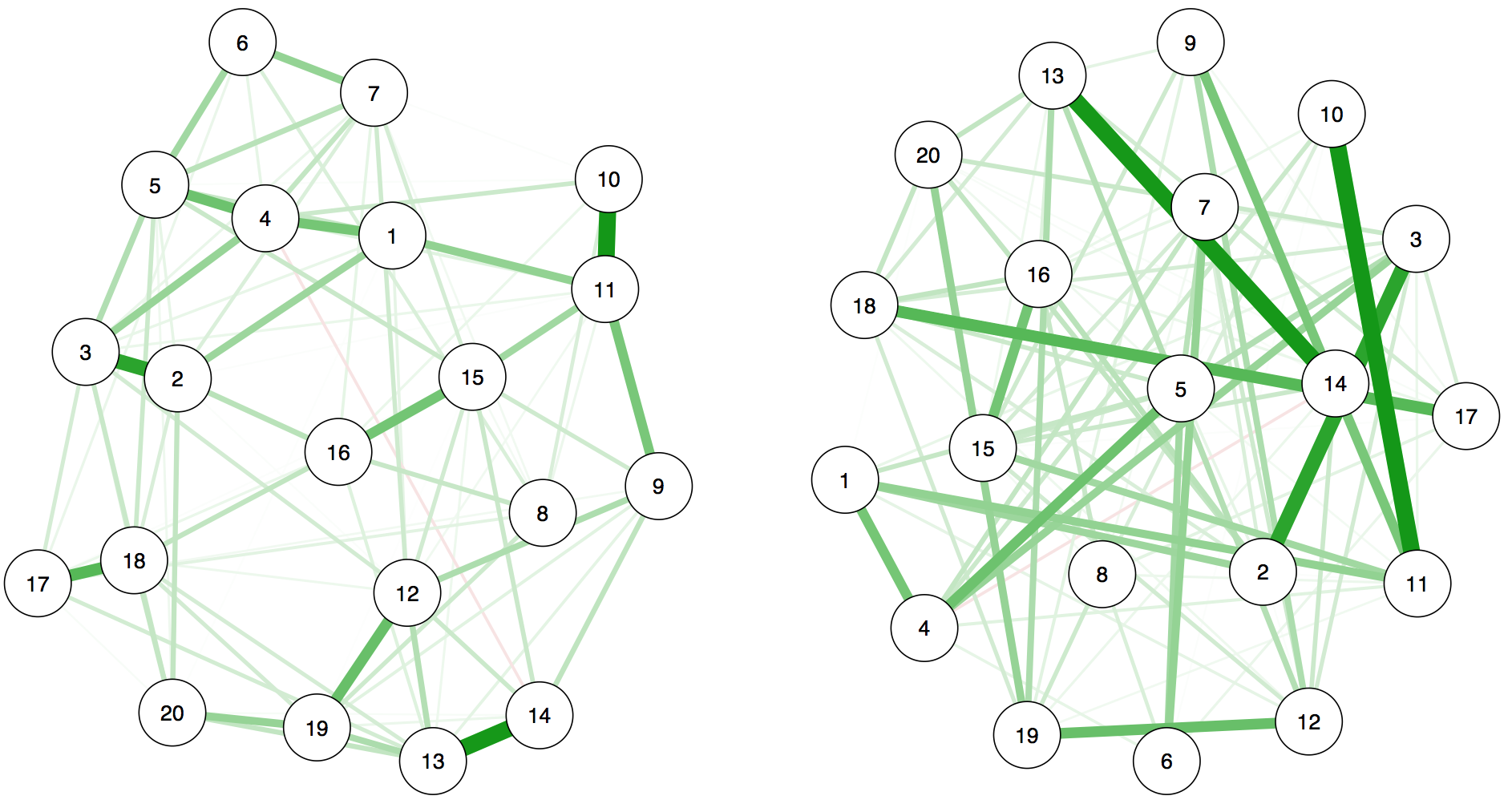
They are exactly identical, and here is the code (download the Rdata file here).
library("qgraph")
library("bootnet")
load("data.Rdata")
pdf("blog1.pdf", width=9.5, height=5.5)
layout(t(1:2))
n1<-estimateNetwork(data, default="EBICglasso")
n2<-estimateNetwork(data, default="EBICglasso")
g1<-plot(n1, layout="spring", cut=0)
g2<-plot(n2, layout="spring", cut=0, repulsion=0.00000001)
dev.off() |
On the right side, I simply changed the repulsion argument so that nodes would be very far apart: all edges are literally the same weight, both in the model result and graphically. This visualization is uninformative, and it is very similar to giving people two correlation matrices where you change columns and rows and then ask them how similar the matrices are. To enable the comparison of rows and columns of two matrices, nodes need to be in the same place in two networks.
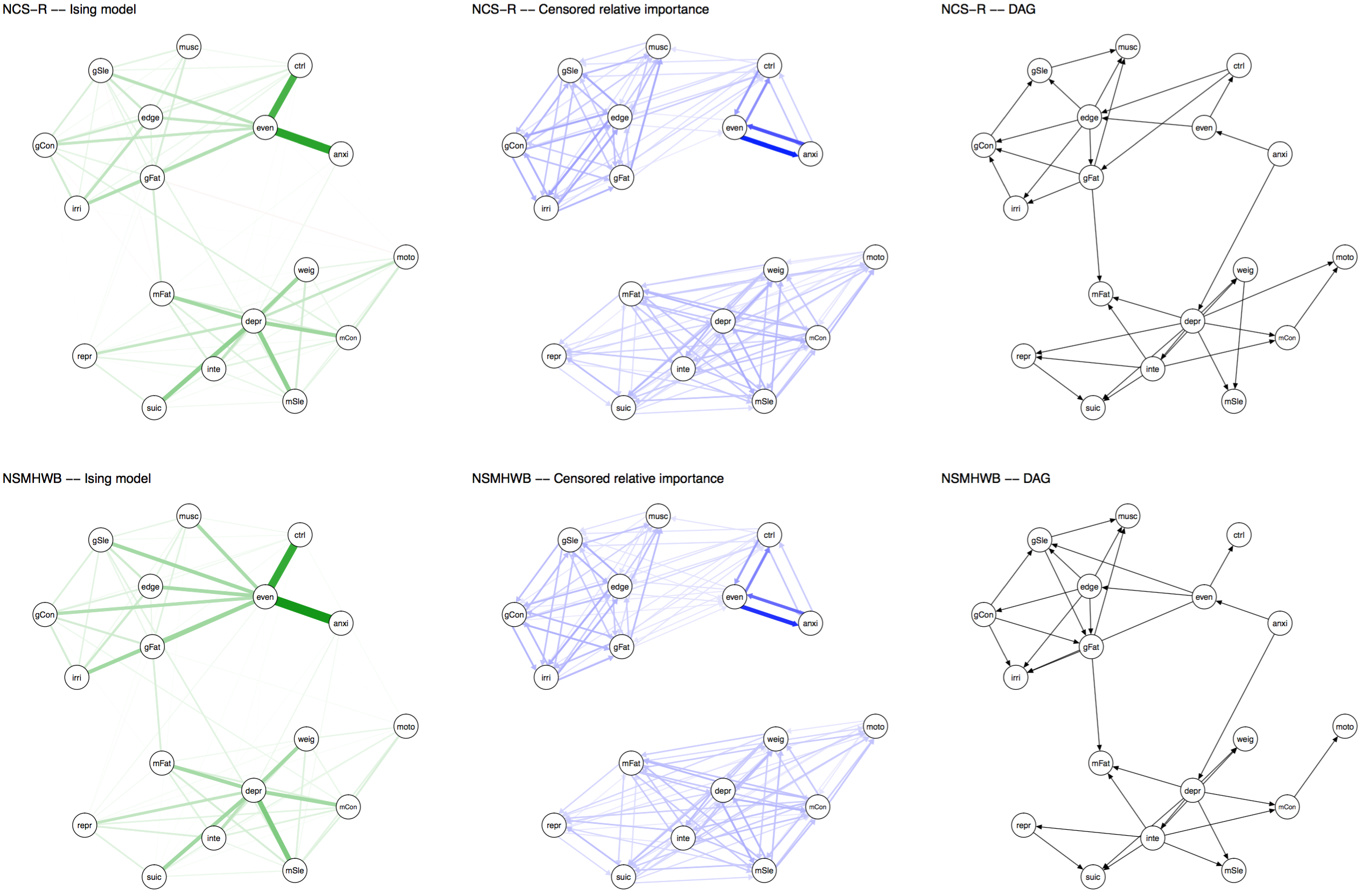
The same holds for network structures. Note that the visual comparison is not very important anyway — we should compare models statistically, not based on visualization, as I’ve highlighted in a previous blog post and in many recent reviews. But FWMK decided to provide graphs in their paper in a way that is uninformative, so it makes sense to post the updated graphs from our commentary here (click the thumbnail for a larger and more legible version of the networks).
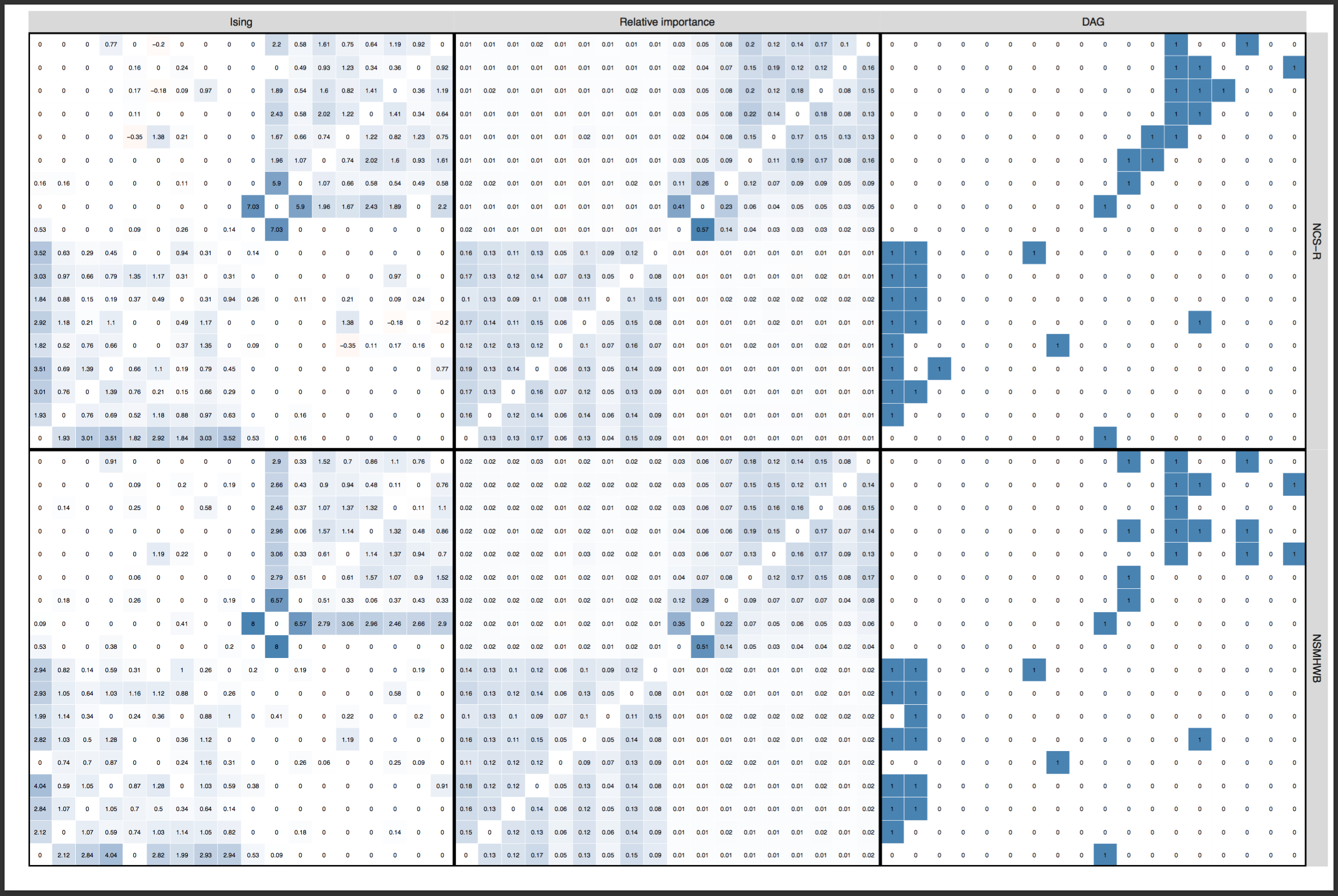
Update November 12: Dr Forbes commented below, strongly pushing back the argument that layouts should be constrained (“ridiculous”, “outrageous”). I am honestly surprised, and had anticipated that we could agree on this point after the explanations above. So I will give this yet another try: below is a visualization (provided by Sacha Epskamp) of the adjacency matrices (in the form of heatmaps) that are used as input for the network graphs, across the two datasets. This is just another way to visualize the edge weights of the networks. Not only is it clear that the conclusion of FWMK that networks do not replicate is not warranted — it also shows why a constrained layout is important, and I could honestly not see anybody argue that we should not constrain the layout of these heat maps to ensure the same edges are in the same rows and columns across 2 datasets. They enable comparison, and do not “obscuring the differences”; constraining the layouts of the network graphs is exactly the same point. Click for full size PDF; reproducible codes and data here (thanks Sacha).
Network estimation versus network inference
The Ising Model was developed in 1917, is very well understood, and has been used in physics, machine learning, artificial intelligence, biometrics, economics, image processing, neural networks, and many other disciplines. Although the implementation to psychology only happened fairly recently16, I find it remarkable that FWMK — after having never worked with the model before, and after fitting it to two datasets — feel confident to conclude that it is “plagued with substantial flaws”. I would not have the confidence to use, for instance, machine learning methodology for the first time, and then write a paper confidently attacking a class of well-established statistical models.
If you read the paper more closely, much of their argument surrounds centrality estimates, and so it is worth mentioning that network estimation and network inference — the interpretation of network topology after you estimated it — are different analytic steps and should not be confused17. And while network methodology is worked out fairly well, the inference is indeed a lot more difficult, and I will return to it at the end of the blog where I hope to find common ground with FWMK. There are definitely problems with network inference, and we need ton of thorough investigations to work these out.
In FWMK’s paper and rebuttal, there are many instances where the authors confuse methodology with the interpretation of methodology, i.e. inference. If you want to criticize the methodology of regression analysis, go ahead and perform a simulation study. If you want to criticize how people interpret regression coefficients, because you do not think these interpretations follow from the regression model, then models are not the problem, but inference. But these two points are very different things, and while the paper by FWMK is clearly focused on the first (see title, abstract, or general scientific summary that is about methodology, not interpretation), the rebuttal pretends it was about interpretation all along, in several sections, but then, again, concludes that “current psychopathology network methodologies are plagued with substantial flaws”.
It does not follow.
Borsboom et al. — our group — published a brief rejoinder to the rebuttal here.
Mistakes are normal and can happen to everybody
I’ve made mistakes, and I think over the course of a scientific career, everybody will. And it is really important to highlight that this is not the issue here. The issue is that a team of authors for the first time used a specific class of psychometric models, made major mistakes in the implementation of these models, drew inferences that do not follow from the results, and then, in their rebuttal, instead of clearly identifying and correcting these mistakes, one-upped their original conclusions with even harsher ones.
Interestingly, when you read their rebuttal, you will notice that FWMK don’t refute any of the points of our rejoinder. Instead, they develop two new arguments: they cite the second commentary that was written on their paper by Steinley et al. numerous times to support their argument, and bring a new argument to the table: PTSD network replicability.
The commentary of Steinley et al. was not actually a commentary on the paper of FWMK, but a critique of the stability of network models, in which the authors propose a new methodology to vet network models. Sacha Epskamp looked into these models and provided a thorough and reproducible refutation of the methodology here18. To summarize, Steinley et al. simulate data from what they suggest to be a proper null model, a random model, but they actually simulate data from a Rasch Model — which is a fully connected network model, not an empty one. So instead of a flat eigenvalue curve where you have no structure in the correlation matrix, they simulated from a model that leads to one very strong first eigenvalue. Their conclusions, therefore, do not follow, because deviations from their null model are not, as they interpret, “indistinguishable from what would be expected by chance”; chance does not lead to a fully connected network or a Rasch model.
I will reply to the second point in the rebuttal of FWMK, PTSD, later.
Common ground
It’s time to move forward, and make the best of this awkward situation. And I’d argue that this shouldn’t be too hard, actually, because FWMK highlight several points in their paper and rebuttal I agree with.
Vetting statistical methodology, and adequate interpretation of statistical parameters, is crucial before drawing substantive (e.g. clinical) inference. FWMK highlight centrality as a problematic parameter that has been thoroughly over-interpreted by some researchers, and I couldn’t agree more. All workshops and lectures on centrality I gave in 2017 contain at least one slide on being careful with interpreting centrality, and here is part of a review I wrote 6 weeks ago:
“My main concern is the terminology and conclusions surrounding centrality. Many previous papers weren’t entirely clear about the potential relevance of central symptoms — maybe the authors could invest a bit more work in this. Playing devil’s advocate here, the most central symptom is likely the most difficult to treat, because after turning it “off” (thinking in terms of the binary Ising Model) it would likely be turned on again due to all the connections. This means that many of the authors’ conclusions regarding treatment do not necessarily follow, and I would be much more careful with clinical implications.”
This is from one of my papers accepted a few days ago:
“It is important to highlight that centrality does not automatically translate to clinical relevance and that highly central symptoms are not automatically viable intervention targets. Suppose a symptom is central because it is the causal endpoint for many pathways in the data: Intervening on such a product of causality would not lead to any changes in the system. Another possibility is that undirected edges imply feedback loops (i.e. A—B comes from A↔B), in which case a highly central symptom such as insomnia would feature many of these loops. This would make it an intervention target that would have a strong effect on the network if it succeeded—but an intervention with a low success probability, because feedback loops that lead back into insomnia would turn the symptom ‘on’ again after we switch it ‘off’ in therapy. A third example is that a symptom with the lowest centrality, unconnected to most other symptoms, might still be one of the most important clinical features. No clinician would disregard suicidal ideation or paranoid delusions as unimportant just because they have low centrality values in a network. Another possibility is that a symptom is indeed highly central and causally impacts on many other nodes in the network, but might be very difficult to target in interventions. As discussed in Robinaugh et al. (Robinaugh et al., 2016), “nodes may vary in the extent to which they are amenable to change” (p. 755). In cognitive behavioral therapy, for example, clinicians usually try to reduce negative emotions indirectly by intervening on cognitions and behavior (Barlow, 2007). Finally, a point we discuss in more detail in the limitations, centrality can be biased in case the shared variance between two nodes does not derive from an interaction, but from measuring the same latent variable.
In sum, centrality is a metric that needs to be interpreted with great care, and in the context of what we know about the sample, the network characteristics, and its elements. If we had to put our money on selecting a clinical feature as an intervention target in the absence of all other clinical information, however, choosing the most central node might be a viable heuristic.”
And it is true that we see in the analysis of the accuracy of centrality that they are often not estimated very reliably, and I have largely stopped looking into analyzing betweenness and closeness centrality in recent months because they often fail to meet minimal criteria for parameter stability.
This brings us to the accuracy of statistical parameters, and I agree with FWMK that this is a crucial topic. When writing up one of my first network papers in 201519, I was very unhappy that I could only obtain an order of centrality values, without knowing whether node A was substantially (or significantly, if you want) more central than node B. So Sacha Epskamp and I sat down for months and tried to find solutions to the problem, and we ended up developing what later became bootnet, a package for testing the accuracy of network parameters. Our tutorial paper on network accuracy20 was published a few months ago and has already gathered about 60 citations — and I wrote a brief blog on the topic here — which means that many researchers adopted the package quickly because they are interested in the accuracy of network parameters to help them draw proper inference. I think that’s a great sign for a field21.
And I have always highlighted that bootnet is definitely not the answer™, but a starting point, e.g. in my network analysis workshops in the sections on stability, and that we need more methodologists pick up critical work. So if FWMK suggest to use split-half reliability for network studies instead of bootnet, I think that’s an interesting complementary approach, and I would like to see some simulation studies to find out how this method performs when we know the true model.
I also agree with FWMK that we need to be vocal about potential challenges and misinterpretations. But I think that many of us have a pretty good track record here. I do that at least once a month when I review network papers, have written several blog posts to safeguard against misinterpretation of network models and parameters (e.g. don’t overinterpret networks visually; don’t interpret coefficients without looking at the accuracy of these coefficients), and have written three papers on the topic. The first paper with Angelique Cramer discusses at length 5 challenges to network theory and methodology, and one of these is replicability22. The second paper — with Sacha Epskamp and Denny Borsboom — is a tutorial on estimating the accuracy of network parameters, which contains a section on replicability. For instance, we state that “[t]he current replication crisis in psychology stresses the crucial importance of obtaining robust results, and we want the emerging field of psychopathological networks to start off on the right foot”23. Third, because we have seen some people misinterpret regularization, Sacha Epskamp and I wrote a tutorial paper in which we explain regularization to applied researchers, and tackle some common misconceptions (e.g. conditioning on sum-scores) and problems24. And I’m obviously not the only person who has been critical here. I urge you to read Sacha Epskamp’s dissertation, which features several outstanding papers. You will find a thorough, careful discussion of network inference, and both the bootnet paper we wrote together, and the discussion of Sacha’s dissertation, deal critically with centrality metrics. Or look at the great critical work by Kirsten Bulteel and colleagues25, or Berend Terluin and colleagues26, or Sinan Guloksuz and colleagues27.
And this goes beyond papers: I also gave talks at numerous conferences urging applied researchers to be careful about interpreting networks, e.g. at APS 201628; and my workshops always include sections on challenges, limitations, and common misconceptions29. I know this is as lot me me me, but I want to clarify that if the topic is interpretation and application, I really hope we can work together on this instead against each other. I am very much interested in this, and have spent the majority of my last 3 years on working on this. I’d like to be an ally, not an adversary.
FWMK also make an interesting point that I hadn’t given much thought before: the overall interpretation of the results of factor models and network models rests on somewhat different parts of the parameter space. For factor models, it is often about the number of factors, and maybe how much variance they can explain in the data, while few researchers would write up in the results section that “factor loading for x1 is substantially larger than factor loading of x2”. We do, however, often see interpretations about strongest edges and most central items in network analysis. This means that we can separate local features from global features of the parameter space. Global features are, for instance, the number of factors vs the number of communities, while local features are the specific factor loadings vs the specific centrality estimates. As I highlighted on Twitter a few days back, global features will replicate similarly badly or well for factor and network models, and local features will also replicate similarly badly or well for both, because the models are mathematically equivalent. The only difference here can be due to the precision of parameters, which is a function of number of parameters we estimate; precision will be somewhat lower for networks, which is one reason network models often use regularization techniques. Apart from that, replicability for both types of models will be the same, and we show that the replicability metric FWMK invented to vet local features of network models performs equally badly for local features of factor models in our commentary. This means that any conclusion FWMK draw about network methodology must hold for factor methodology, and this is not an opinion, but based on mathematical proofs.
PTSD replicability
The rebuttal of FWMK also contains a table on PTSD network papers.
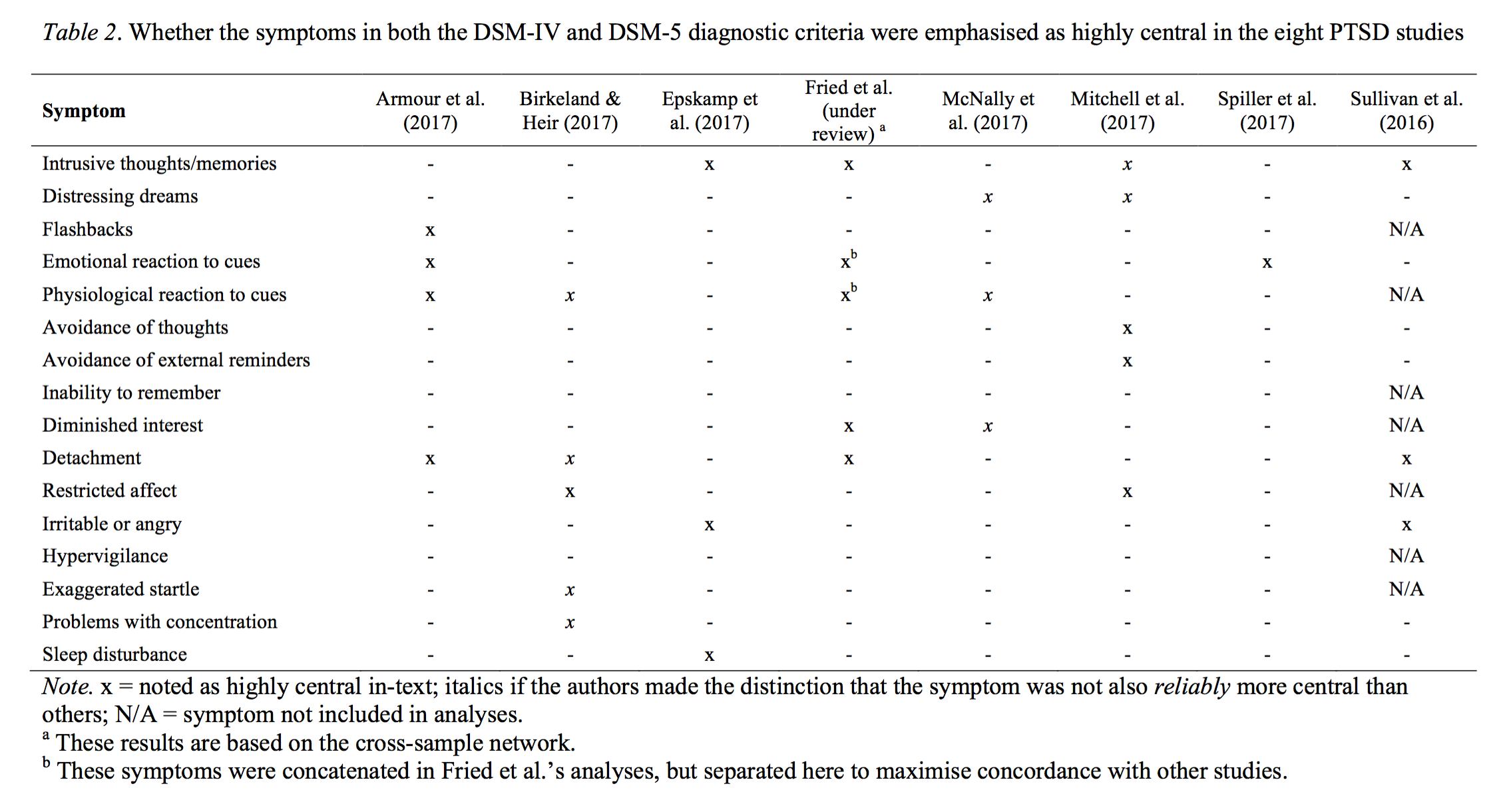
The table, while somewhat incomplete (you can find a full list of all PTSD network papers I know of here), clearly highlights the substantial heterogeneity in the PTSD network literature, which is the very reason that I started a large interdisciplinary multisite PTSD network replicability project end of 2015. In our paper that was accepted in Clinical Psychological Science just this week — I have written up the main results in a blog post here — we estimate networks in 4 clinical datasets of patients receiving treatment for PTSD, which I find to be more informative than community data when it comes to network structures. The data also have no skip problems, and thus circumvent the majority of the problems inherent in the data of FWMK.
Conclusions
There are a ton of other challenges that lie ahead, many of which are not only statistical, but conceptual (again, the methodology really is just fine). If you are interested, I mention numerous papers above that I link to in footnotes. To tackle these issues, it is crucial that both applied and methodological researchers pick up network methodology, distinguish clearly between network theory and network methodology, advance network theory, vet network methodology, think clearly about interpretation, and voice their criticisms.
If possible, however, I would like to do that in cooperation with others, rather than as adversaries. Working together is a lot more fun than writing critical commentaries and blogs. This is consistent with the personal take of Sacha Epskamp on the topic, who published a post-publication review about the target paper discussed here on PubPeer. Like me, Sacha concludes that there are numerous important issues that need to be explored further in the future, highlights critical work he and colleagues conducted — but also points out that the conclusions by FWMK do not follow from the evidence they present.
Update December 04 2017
Sacha Epskamp posted his personal comments on the paper as post-publication review on pubpeer.com.
Footnotes
- Or maybe it does, in any case I wrote it and I’m responsible for the content. Sacha Epskamp, whom I sent the draft prior to publishing, said he’d want to sign it, so I mention his name here specifically.
- The authors did share their code, and were very responsive to our emails, which is what enabled us to identify the problems.
- March 4th 2019: I had uploaded the 3 different final versions here originally, but just found out that Google Scholar indexed them, which leads to confusion about what the final version of the paper is. I have therefore removed the links to prior versions, which are from now on available per request
- FWMK brought to my attention that 2 editors looked over the paper after these changes, however.
- In their rebuttal, the authors state that readers should consider the relative importance networks from our reanalysis instead of those provided in their original paper.
- We obviously also teach factor models, and regressions, and Bayesian statistics, and t-tests … as you can see, we have extremely severe conflicts here.
- I mention simulations several times in this blog; mathematical derivations are preferred, of course, but since I can hardly spell the word ‘math’, I prefer simulation studies because I understand what they can and cannot do.
- There are also k threshold parameters, but let’s ignore them here to keep things simple.
- Correlations don’t work for DAGs, see our commentary.
- FWMK concluded 74.2% originally, but due to an error in their implementation of the methods, they concede in their rebuttal that our estimate is the correct one
- This test has been developed by Claudia van Borkulo and colleagues for the Ising Model, the network most commonly used so far in the literature. This test is very conservative and tests whether all edges across two networks are exactly identical; van Borkulo, C. D., Boschloo, L., Kossakowski, J. J., Tio, P., Schoevers, R. A., Borsboom, D., … Boschloo, L. (2017). Comparing network structures on three aspects. http://doi.org/10.13140/RG.2.2.29455.38569
- Note that I wouldn’t argue listwise deletion is the way to go here, but the differences between the two methods of dealing with missing data are astonishing.
- See footnotes 6-10 in Cramer, A. O. J., Waldorp, L. J., van der Maas, H. L. J., & Borsboom, D. (2010). Comorbidity: a network perspective. The Behavioral and Brain Sciences, 33(2–3), 137–50. http://doi.org/10.1017/S0140525X09991567
- e.g. Boschloo, L., van Borkulo, C. D., Rhemtulla, M., Keyes, K. M., Borsboom, D., & Schoevers, R. A. (2015). The Network Structure of Symptoms of the Diagnostic and Statistical Manual of Mental Disorders. Plos One, 10(9), e0137621. http://doi.org/10.1371/journal.pone.0137621; the paper contains contains a section entitled “Sensitivity analyses (dealing with skip-related missingness)”, in Boschloo et al. state: “As implied by the skip logic, the skip-related missing values on the non-screening questions were imputed with zeros, indicating absence. This imputation strategy may have artificially induced strong connections within diagnoses and weak or absent connections between diagnoses”.
- Borsboom, D., & Cramer, A. O. J. (2013). Network analysis: an integrative approach to the structure of psychopathology. Annual Review of Clinical Psychology, 9, 91–121. http://doi.org/10.1146/annurev-clinpsy-050212-185608
- van Borkulo, C. D., Borsboom, D., Epskamp, S., Blanken, T. F., Boschloo, L., Schoevers, R. A., & Waldorp, L. J. (2014). A new method for constructing networks from binary data. Scientific Reports, 4(5918), 1–10. http://doi.org/10.1038/srep05918
- See e.g. this introductory tutorial: Epskamp, S., Borsboom, D., & Fried, E. I. (2017). Estimating Psychological Networks and their Accuracy: A Tutorial Paper. Behavior Research Methods, 1–34. http://doi.org/10.3758/s13428-017-0862-1
- Preprint updated March 9th 2018
- Fried, E. I., Epskamp, S., Nesse, R. M., Tuerlinckx, F., & Borsboom, D. (2016). What are “good” depression symptoms? Comparing the centrality of DSM and non-DSM symptoms of depression in a network analysis. Journal of Affective Disorders, 189, 314–320. http://doi.org/10.1016/j.jad.2015.09.005.
- Epskamp, S., Borsboom, D., & Fried, E. I. (2017). Estimating Psychological Networks and their Accuracy: A Tutorial Paper. Behavior Research Methods, 1–34. http://doi.org/10.3758/s13428-017-0862-1
- You know who didn’t use bootnet for testing stability of their analysis? FMWK ;) …
- Fried, E. I., & Cramer, A. O. J. (2017). Moving forward: challenges and directions for psychopathological network theory and methodology. Perspectives on Psychological Science, 1–22. http://doi.org/10.1177/1745691617705892]
- Epskamp, S., Borsboom, D., & Fried, E. I. (2017). Estimating Psychological Networks and their Accuracy: A Tutorial Paper. Behavior Research Methods, 1–34. http://doi.org/10.3758/s13428-017-0862-1
- Epskamp, S., & Fried, E. I. (2017). A Tutorial on Regularized Partial Correlation Networks. Accepted in Psychological Methods. https://arxiv.org/abs/1607.01367
- Bulteel, K., Tuerlinckx, F., Brose, A., & Ceulemans, E. (2016). Using raw VAR regression coefficients to build networks can be misleading. Multivariate Behavioral Research. See also this really cool preprint!
- Terluin, B., de Boer, M. R., & de Vet, H. C. W. (2016). Differences in Connection Strength between Mental Symptoms Might Be Explained by Differences in Variance: Reanalysis of Network Data Did Not Confirm Staging. Plos One, 11(11), e0155205. http://doi.org/10.1371/journal.pone.0155205; I also wrote a blog summarizing the paper.
- Guloksuz, S., Pries, L., & Van Os, J. (2017). Application of network methods for understanding mental disorders: Pitfalls and promise. Psychological Medicine, 1-10. doi:10.1017/S0033291717001350.
- “How to increase robustness and replicability in psychopathological network research”.
- Here all materials for the September 2017 workshop I gave in Madrid.

Hi Eiko,
My impression of this blog post and your comments on social media is that you are misrepresenting the process and content of the papers, as well as questioning the competence and integrity of me and my co-authors. This is disappointing. Such an approach distracts from the key issues and only hinders a constructive and collaborative way forward.
Surprisingly, this blog post restates many of the points in your published commentary while appearing to ignore our rejoinders: Nearly half of the text of our reply falls under the heading “Borsboom et al.’s main criticisms of our article” (p. 1012-1013) with each paragraph addressing a key point. Further, Table S1 “Brief responses to less central points from Borsboom et al.’s (this issue) commentary” also addresses less central points made in the commentary. I would encourage anyone interested in this exchange to read the four peer-reviewed papers (our original paper, the two commentaries, and the reply), rather than social media/blog posts. Please feel free to contact me for copies of the papers if you do not have access.
Having said that, for those readers who may have concerns based on the points that are made in this post, I offer four additional comments about some of the central points:
1.) Both this blog post and the earlier commentary suggest that our publication is riddled with serious errors/flaws/problems that undermine our results. We addressed the specific criticisms in our reply. The only analytic decision that I think could be labelled an error in our publication was an unintended consequence of the censoring rule we used for the relative importance networks: Borsboom et al. pointed out in their commentary that our censoring rule—intended to isolate the edges with high relative importance—unintentionally removed both edges in a pair when they had nearly identical edge weights. We acknowledged and addressed this point in our response. Notably, our conclusions hold independently of the censored relative importance networks (we also analysed uncensored relimps); independently of all relative importance networks (comparing only Ising models and DAGs, which are better suited to analysing binary data); and even independently of all of our results (based on a review of the extant PTSD literature using “state-of-the-art” analyses).
2.) The review process is also misrepresented. For example, you imply that we essentially re-wrote our article after acceptance. After being surprised by the psychosystem group’s strong emotional response to our initial pre-print, at the copy-editing stage we tried to read the paper from your perspective and revise any language that may be perceived as antagonistic (e.g., removing generalisations about how researchers apply the methods, and toning down the title). We subsequently discovered and fixed the error in the DAGs a few days after those copy edits were completed. Two editors from the journal reviewed the corrections, agreeing they did not change our conclusions, and allowed us to correct the copy-edited manuscript before publication. As an aside, it is remarkable that it hardly made a difference to the replicability of the DAGs whether our results were accurate (analysing the raw data as input) or nonsense (analysing the correlation matrices as input).
3.) This blog post also emphasises the point that our paper cannot address the replicability of psychopathology networks by comparing networks fit to two datasets because each dataset may have a different underlying model. You said above “Unfortunately, the authors did not address this point in their rebuttal.”. We did (see p. 1012). One key point that we may not have emphasised enough in our reply is that only one of the twenty-one sets of analyses in our paper was based on comparing the two large (n > 8800) community datasets. The remaining 95% of the analyses are based on randomly splitting the datasets in half 20 times (i.e., creating 10 pairs of random split-halves within each dataset) and comparing the estimated networks in each pair—which would not be expected to have different underlying structures. Such real-world data evaluations are just as important as simulation studies for understanding how data perform; simulation studies rarely achieve comprehensive evaluation of all the data features that might be of interest, and simulated data often fail to include the types of real-world variations that can lead to different and poor model performance in applied settings.
4.) Finally, I note that you state we are “attacking a class of well-established statistical models”. Our paper is not criticising the application of network methods in general, but their application in symptom-level psychopathology data specifically. I could not find examples where we referred to “network methods” in general, instead of the application of these methods in psychopathology, and I re-iterate that we are not questioning whether Ising models can reliably and validly be applied in physics, for example. The supplement to our original paper explains why the reliance of these models on conditionally independent relationships means that poor replicability is pretty much inescapable in psychopathology symptom networks. While extraordinarily large sample sizes (e.g., hundreds of thousands) or very precise measurement could be a solution, we are unfortunately limited by the realities of the data we analyse in our field.
These are just a few key points. I believe the large majority of the condemnations in this post and on social media are either invalid or untrue. If anyone feels there are important points that we have not addressed (e.g., why it is ridiculous to criticise our use of the Fruchterman and Reingold algorithm to visualise the networks) please don’t hesitate to reach out, as we’d be happy to discuss. We’ll also write a brief comment on the psychosystems blog post, but I personally will refrain from any further public debate or comment on points that have already been addressed. We again strongly encourage anyone seeing this to carefully read the peer-reviewed research in question.
Miri (on behalf of FWMK)
Hi Miri,
Thanks for chiming in. I am happy to continue the discussion as long as we challenge each other to think differently in a civil way, and don’t just repeat the same arguments.
I’ll start with two general points and then address your 4 points with the corresponding numbers.
A) PhD students and more senior researchers in our department, including well-established psychometricians and mathematical psychologists, have worked years to vet network methodology thoroughly, and have published numerous peer-reviewed methodological papers, including many simulation studies, in well-respected methodological journals such as Psychological Methods, Psychometrika, Behavioral Research Methods, and Scientific Reports. They have also spent years of work developing software (*none* of the credit here goes to me, I’m an applied researcher).
Your paper, on the other hand, published in a clinical journal, fit network models to 2 datasets, included no simulations or mathematical proofs, and concluded “Network models are plagued with substantial flaws”. So, yes, as expected, you get pushback by methodologists who have worked hard on developing and vetting these methods, and who understand them well, because your evidence does not justify your conclusions.
B) You claim that the above blog ignores your rebuttal. That is untrue, I discuss the 2 new points you raised in your rebuttal. And I agree with you—and have stated that clearly multiple times—that readers should read your rebuttal and make their own sense of the arguments you provide, and link to the rebuttal. And to remind you, the goal of the blog was not to summarize all arguments, and I say this clearly, the goal was to tell my personal story (that does not necessarily reflect those of other authors involved) of a very weird publication process that, in my opinion, was more than suboptimal (I think all authors involved on all sides agree on this), about a paper that I find very questionable for the reasons I have layed out. I have cited you rebuttal, and encouraged readers to look into it.
1) “Both this blog post and the earlier commentary suggest that our publication is riddled with serious errors/flaws/problems”.
Again, this is flatout wrong: I do not claim your paper is “riddled with” anything. I wrote, quote: “there is at least one known serious error, and a number of major problems.” We agree that there is one error, and while you don’t think it’s a “major problem” fitting linear regression to binary data that would require a logistic regression, it seems pretty well established that this is not really among the things to do.
“Notably, our conclusions hold independently of the censored relative importance networks (we also analysed uncensored relimps); independently of all relative importance networks (comparing only Ising models and DAGs, which are better suited to analysing binary data); and even independently of all of our results (based on a review of the extant PTSD literature using “state-of-the-art” analyses).”
It simply doesn’t follow Miri. Imagine the most extreme case: I make up data completely to make a claim that variables x and y are highly correlated, and write a paper about it. Now people find out I made the data up, and point this out in a commentary. Now I respond with another paper where I actually look into new data, and find x is highly correlated with y (I disagree with some of your PTSD conclusions, but let’s just assume here it’s all correct for the sake of the argument). That doesn’t mean the commentary was incorrect, nor that my original paper was correct.
We never wrote a commentary saying x and y are uncorrelated. We said: your paper does not allow for the conclusion that x and y are highly correlated because of all the issues I list above. Unlike the sweeping generalizations you draw from fitting models to 2 datasets in the paper, we were very clear that our conclusions only pertain to the 2 datasets we analyzed.
2) You claim I misrepresent the review process in two parts. Both claims are wrong.
First, you state I claimed you “essentially re-wrote our article after acceptance”. I never said that. I pointed out, factually correct: “In total, FWMK fixed the DAG errors, rewrote parts of the paper, changed the title (“…”) and the results, but left the discussion and conclusions untouched.” So, no, I did not misrepresent that.
Second, you say that 2 editors looked at your changes you performed after the final accepted version of the manuscript, and agreed with them. I never claimed anything to the contrary. I literally wrote: “The paper was not peer-reviewed after the changes.” So, no, I did not misrepresent that either, because peer-review is (to my knowledge) not defined as editors looking over a paper. But I will edit the blog to make sure this is mentioned. Incidentally, it’s good to hear that Abnormal had editors check your changed version of the paper, although I wasn’t aware that they have methods experts who can vet Directed Acylic Graphs analyses.
3) I am confused that after a paper, a commentary, a rebuttal, and my blog post, you still mix up model stability with model replicability. Please read title, abstract, and scientific summary of your own paper: you talk about replicability, defined as the question to which degree a finding replicates in a different dataset (out of sample prediction). Model stability is the question about how accurate parameter of a given model are estimated. Both are important, and model stability is a prerequisite for replicability — but they are very different things.
Now, you did perform split-half analyses, and I liked that, and said so in the blog post above. I think that’s a great idea, and I never criticized you for these, neither did Borsboom et al. I personally think it is a neat idea, well established, and (after some simulations, because sample size impacts on network structure due to regularization, so split half will not work in small samples) I would consider adding that to the network toolbox. But split-half stability is not the same as replicability (which is what you claim to test in the paper). I disagree on your statement on simulation studies, but I’m ok to just disagree here and let it go.
4) “Our paper is not criticising the application of network methods in general, but their application in symptom-level psychopathology data specifically. […] The supplement to our original paper explains why the reliance of these models on conditionally independent relationships means that poor replicability is pretty much inescapable in psychopathology symptom networks.”
I seem to be missing crucial information somewhere between (A) the fact that you have a diagram of a partial correlation in our supplementary materials and (B) your claim that partial correlations have poor replicability. I don’t think one follows from the other, and I also don’t understand why this should be specifically so for symptom data (and not intelligence or personality or emotion data).
In any case, when it comes to replicability, we go into the point in the commentary in detail, and I said in the beginning that I don’t think this is a useful exercise if we repeat ourselves. So I will just say that you invented and used a metric of replicability that has the following features:
– The metric is neither vetted nor validated; it has not been published anywhere else; you have never used it before, neither have other people (as far as I know). You also don’t show that this metric actually works, for which you would need to simulate from the same and two different models, and show what the sensitivity and specificity of the metric is to actually pick that up.
– The metric shows that 2 network models do not replicate although literally every other single test shows they are identical or nearly identical (both in your paper and in our re-analysis), and although the only validated psychometric test—the network comparison test—shows that they are exactly identical (for which there are simulation studies on which the test was validated, by the way).
– Your metric shows that factor models replicate even worse than network models, even if we simulate them from the same true model (we did the simulation work for your test; it doesn’t work). It is stunning that you keep repeating the same argument after we showed that this is the case; see our commentary, search for “MIRT”.
—-
To conclude, I disagree that I (or my arguments) are “ridiculous” — I put a lot of effort into making all arguments broadly accessible, including e.g. data and R syntax to explain the visualization problems. In any case, you will not find that I made similar claims about you or anyone else on FWMK on social media, this blog, or elsewhere. And I think it’s important to criticize each other’s work to move things forward. And, as I said in the blog post, I’d much rather do that together (I am as skeptical as you about things like centrality) than as adversaries.
Cheers
Eiko
(updated 11/11/17 0.27am local time)
Hi Eiko,
I understand that your research group is heavily invested in these methods, and therefore criticisms of their performance would motivate you to respond strongly. However, I’m not sure you are fully aware of the messages implied by the tone and content of your blog posts. It seems to me that this interaction is not heading in a constructive direction. The new points you have raised here are addressed in our reply, and I agree with you it is not a useful exercise to repeat ourselves.
One final point: I did not at all mean to suggest that you are ridiculous, and I’m sorry that my comment could be read that way. My point was that it is outrageous to criticise our use of the Fruchterman and Reingold algorithm to plot the networks in our paper, and I should have worded it more thoughtfully.
Miri
Hi Miri,
Chiming in on this conversation, as you refer to our research group of which I am part of. I don’t think any of us is “heavily invested” in these methods, as we do ample of work on criticising network interpretations (https://doi.org/10.1371/journal.pone.0179891), challenges to network analysis (https://osf.io/bnekp/), stability and accuracy (https://doi.org/10.3758/s13428-017-0862-1), incorporating measurement invariance (https://doi.org/10.1007/s11336-017-9557-x), developing methods for comparing two groups (https://www.researchgate.net/publication/314750838_Comparing_network_structures_on_three_aspects_A_permutation_test) , generalizing to time-series data (https://arxiv.org/abs/1609.04156), and replicability (https://osf.io/2t7qp/). If anything, your work reads as an advertisement for my work. I can’t speak for my colleagues, but I criticise your work not because of being “invested”, but simply because it does not paint a correct picture. I’d also like to note that I think the tone of Eiko’s blog is far better than the tone and content of your published work (e.g., “the current psychopathology network methodologies are plagued with substantial flaws”).
The reason I am writing this reply, though, is because of your comment towards the use of Fruchterman-Reingold. I don’t see Eiko or anyone criticising you for using Fruchterman-Reingold, which we use all the time. The criticism is directed at you interpreting the node placement and expecting node placement to replicate. Fruchterman-Reingold is a chaotic algorithm, and a tiny change in the input may result in an entirely different node placement. This is the point Eiko describes in detail in this blog post. It is also important and the reason why you should constrain layouts to be equal. After keeping the layouts constrained (still using Fruchterman-Reingold) it is very clear that the estimated networks are very similar indeed. Of course, the Fruchterman-Reingold algorithm is only meant as a visual tool and should not be interpreted in any other way: the node placement is in no way connected to, e.g., centrality. I, at least, always make this very clear in teaching on these methods.
It seems as if you think we unfairly attack you for doing things we do too, given your replies to the zero-imputation and Fruchterman-Reingold algorithm. These are not meant as low blows, but merely because these are important points for consideration. I hope you will not take or make this discussion more personal than it needs to be.
Hi Sacha,
I was referring to Eiko’s point A) above, which I understood to be a strong statement of your group’s investment in the methods.
I’ll comment briefly on the Fruchterman-Reingold algorithm point, given I gave it as an example of something we didn’t think was particularly important to address directly. As you say, this algorithm is used almost invariably in the literature to plot psychopathology networks, so we did the same. We did not presuppose that any specific network characteristics would replicate (or not replicate), but were testing this as a research question. If you revisit our paper, you will see that rather than “interpreting the node placement and expecting node placement to replicate”, we explicitly cautioned researchers against doing this (footnote 4 and p. 982).
Constraining the layouts to be equal artificially creates an impression of greater similarity between networks, obscuring the differences between networks that are evident in our metrics. This was a key point in our reply: The metrics used in the Borsboom et al. commentary/re-analysis lack sensitivity to evidence for poor replicability.
I don’t see your comment above as a “low blow” or personal attack, and please note that I have consistently focused my contributions to this discussion on the research without attacking any of you personally. Please note too that I intend to step back from further discussion until I return from leave in early 2018.
Miri
Regarding my statement A), Miri, you just need to read it to the end. I say clearly why you received pushback.
“A) PhD students and more senior researchers in our department, including well-established psychometricians and mathematical psychologists, have worked years to vet network methodology thoroughly, and have published numerous peer-reviewed methodological papers, including many simulation studies, in well-respected methodological journals such as Psychological Methods, Psychometrika, Behavioral Research Methods, and Scientific Reports. They have also spent years of work developing software (*none* of the credit here goes to me, I’m an applied researcher).
Your paper, on the other hand, published in a clinical journal, fit network models to 2 datasets, included no simulations or mathematical proofs, and concluded “Network models are plagued with substantial flaws”. So, yes, as expected, you get pushback by methodologists who have worked hard on developing and vetting these methods, and who understand them well, because your evidence does not justify your conclusions.”
Regarding the Fruchterman-Reingold, I showed an example in the blog where networks look very different although they are exactly the same, showing that unequal layouts are misleading and uninformative. You called this argument first “ridiculous”, and then, later “outrageous”, but that will not get you around this fact. The same holds for correlation matrices. When we compare 2 correlation matrices, we will constrain the layout, meaning we have the same variables in the same columns and rows across 2 matrixes to facilitate comparison. You *can* argue that this “artificially creates an impression of greater similarity between matrices” — this is the same argument you use for networks — but I don’t think it’s a reasonable argument. And I don’t think anybody else will think it is.
Hi Eiko,
To clarify, we didn’t do anything to cause unequal layouts in our networks (e.g., changing the repulsion argument/scrambling rows and columns in matrices like you do in the examples in your post). We applied the default Fruchterman-Reingold algorithm to all networks, just as it’s applied in the extant literature. My point is that criticising this analytic decision—e.g., suggesting that the resulting layouts are “misleading and uninformative”—runs directly counter to the bulk of the extant literature.
Also, I haven’t suggested that constraining the layouts to be equal is either “ridiculous” or “outrageous” (per your misattributions in the updated blog post), but that it creates an artificial impression of similarity between them.
Miri
1) Miri: “We applied the default Fruchterman-Reingold algorithm to all networks, just as it’s applied in the extant literature […] Criticising this analytic decision […] runs directly counter to the bulk of the extant literature..
There is no “bulk of the extant literature”, because very few papers have investigated network structures across multiple datasets — and the whole point is about papers that do so. You cannot constrain layout equal when you only have one network. Also, plotting networks is not an analytic decision, it’s a data visualization decision.
How do things look like in the extant literature? Like this, for instance. Notice the constrained layouts to facilitate interpretation.
(Rhemtulla, M., Fried, E. I., Aggen, S. H., Tuerlinckx, F., Kendler, K. S., & Borsboom, D. (2016). Network analysis of substance abuse and dependence symptoms. Drug and Alcohol Dependence, 161, 230–237. http://doi.org/10.1016/j.drugalcdep.2016.02.005)
Sacha explained the point clearly in his recent comment why this is important:
“The reason I am writing this reply, though, is because of your comment towards the use of Fruchterman-Reingold. I don’t see Eiko or anyone criticising you for using Fruchterman-Reingold, which we use all the time. The criticism is directed at you interpreting the node placement and expecting node placement to replicate. Fruchterman-Reingold is a chaotic algorithm, and a tiny change in the input may result in an entirely different node placement. This is the point Eiko describes in detail in this blog post. It is also important and the reason why you should constrain layouts to be equal.
2) Miri: “I haven’t suggested that constraining the layouts to be equal is either ‘ridiculous’ or ‘outrageous’ (per your misattributions in the updated blog post)”
Blog: “Dr Forbes commented below, strongly pushing back the argument that layouts should be constrained”
Quotes Miri:
– “If anyone feels there are important points that we have not addressed (e.g., why it is ridiculous to criticise our use of the Fruchterman and Reingold algorithm to visualise the networks)”
– “My point was that it is outrageous to criticise our use of the Fruchterman and Reingold algorithm”
Dear Miri,
Your statements regarding the Fruchterman-Reingold algorithm create the impression that you don’t really understand how this algorithm works. As Sacha already mentioned the Fruchterman-Reingold algorithm is a chaotic algorithm, in which tiny differences in the network structure lead to huge differences in the layout. The following R-code is an illustration of this:
library(Matrix)
library(qgraph)
set.seed(1)
graph <- matrix(rnorm(100), 10, 10)
diag(graph) <- 0
graph <- forceSymmetric(graph)
qgraph(graph, layout = 'spring')
graph2 <- graph
graph2[1,2]<-graph2[2,1]<-graph[1,2]-.00000000001
layout(matrix(1:2,1,2))
qgraph(graph, layout = 'spring')
qgraph(graph2, layout = 'spring')
qgraph(graph, layout = 'circle')
qgraph(graph2, layout = 'circle')
When you run this code, you'll see that two network that are virtually identical (the only difference being that the edge from node 1 to node 2 is 0.00000000001 smaller in the second network) are plotted quite differently. When you simply plot the networks in a circular layout, it is much easier to tell that these networks are basically identical (I really don't see why plotting the networks in such a way would "create[s] an impression of greater similarity between networks" as you stated).
Jonas
Hi Jonas,
You may have misunderstood my statements about the algorithm. Your example is a better illustration of the point Eiko is trying to make in the Visualization section above because it shows that trivial differences in the underlying data can effect big changes in the resulting layouts (whereas Eiko’s example shows that different layouts of the same data look different). This chaotic-ness of the F-R algorithm is central to my point: Applying the default algorithm—as we did in our analyses to mirror current practices in the psychopathology network literature—results in very different layouts; forcing these layouts to be equal—as Borsboom et al. did in their commentary—overrides the chaotic nature of the algorithm to create an artificial impression of similarity. Plotting the networks in a circle is a fine alternative to overcome the noise manifested by the F-R algorithm, but doesn’t address any of the fundamental issues raised in the peer-reviewed exchange.
Miri
Hi Miri,
1. You state that “plotting the networks in a circle is a fine alternative to overcome the noise manifested by the F-R algorithm”. Noise is really not the correct term in this context. You can argue that the estimation of networks is affected by noise, but the F-R algorithm simply takes the network you feed it and plots it in the way that it is supposed to do. Important to note here is also that the F-R algorithm was not created for plotting several networks to compare them. As I read your paper, your aim was to investigate whether networks are similar or not. Plotting two networks with the F-R algorithm is simply a poor choice if you want to do this, because you make it virtually impossible for the reader to see to what extent the networks differ.
2. You state that “forcing these layouts to be equal… create an artificial impression of similarity”. First of all, similarity in what sense? Do you mean similar in their layout? If so, your statement is correct, but also extremely trivial, because everyone who has the slightest idea of the working of the F-R algorithm knows that the layout between any two networks won’t be similar. Or do you mean similarity of the network itself? If so, this is not correct, because plotting two networks with the F-R algorithm inflates dissimilarity between networks much more than plotting the networks with the same layout inflates similarity.
3. You state that “plotting the networks in a circle is a fine alternative… but doesn’t address any of the fundamental issues raised in the peer-reviewed exchange”. First of all, I didn’t claim that it would, but my point is that when you talk about the F-R algorithm both in the “peer-reviewed exchange” and in the comments here you very much create the impression that you expect the F-R algorithm to perform in a way that it is not supposed to. For example, in footnote 4 of your original article on “replicability of psychopathology networks” you state that “the placement of nodes and their proximity to each other was unreliable” and you don’t qualify this statement by mentioning that we can’t expect the F-R algorithm to do that.
In sum, I still very much have the impression that you expected the F-R algorithm to perform in a way that it is simply not supposed to. I puzzles me that you don’t see this as an issue and don’t simply acknowledge that it wasn’t the best idea to plot the networks in the way you did.
Jonas
It is a fair argument that constraining layouts may visually inflate similarity. I have played around with semi-constraining layouts to get around this but it is not trivial. However, that doesn’t negate the point that non constraining the layout very strongly inflates dissimilarity. As shown with Jonas’ and Eiko’s codes, it is very hard to visually judge those networks to be about identical if the node placement is different. That argument is not “ridiculous” or “outrageous”.
I see Eiko now uploaded the heatmaps I made as well in the blog post which offers a third visualization of the networks.
Hi Jonas and Sacha,
I’ll try once again to clarify two points that have been repeatedly misunderstood here:
1) We didn’t do anything tricky with our network layouts. The default F-R algorithm has been used almost without exception in the literature to plot psychopathology networks, so we did the same. Notably, we did not use the algorithm “to show how different [the networks] are” or to compare networks (see “Comparing the networks” p. 974-975); and we did not interpret node placement or expect the layouts to be replicable or reliable. We simply plotted the networks as they are plotted in the literature.
2) Using the default F-R algorithm is so overwhelmingly the norm in the literature that I made the point it is “ridiculous”/”outrageous” to suggest we shouldn’t have used it. My use of these evidently controversial words predated all of the subsequent discussion of the algorithm, so did not refer to any specific argument as has been repeatedly misrepresented here. Further, I did not say that it is ridiculous/outrageous to constrain the layouts to be equal (also repeatedly misattributed above) but only highlighted that doing so was just one more way that the similarities between networks were emphasised over the differences by Borsboom et al.
Regardless, the F-R algorithm is largely irrelevant to the key issues in the papers. This discussion is frustratingly unproductive, but I’ve felt compelled to respond when our work is misrepresented and when my competence and integrity are publicly questioned. I’ll renew my efforts to withdraw from this interaction now, and would again encourage anyone interested in the research to read the peer-reviewed exchange.
Miri
P.S. If you’re reading this blog because the peer-reviewed exchange is very long, some sections of our target article that cover key issues include:
The first paragraph of “Conditionally independent networks” (p. 970)
“The utility of psychopathology networks relies on generalisability and replicability” (p. 971-972)
The first paragraph of the discussion (p. 981)
“Why might conditional independence networks have such poor replicability?” (p. 984-985)
Appendix S1
And from our reply to the commentaries:
Table 1 and paragraph 2 on p. 1013
“Evidence of poor replicability from the psychopathology network literature” (p. 1013-1014)
The first paragraph of “The future of psychopathology networks” (p. 1014)
Miri: “Using the default F-R algorithm is so overwhelmingly the norm in the literature that I made the point it is “ridiculous”/”outrageous” to suggest we shouldn’t have used it.”
Your rebuttal remains virtually orthogonal to the argument raised. The point was about constrained vs unconstrained layouts, not about using or not using F-R. And the three different reproducible examples by Jonas, Sacha, and me show that point convincingly (e.g., even identical networks look wildly different using unconstrained layouts).
Miri: “We simply plotted the networks as they are plotted in the literature.”
Not true. The standard in the literature that plot one network each is using F-R, which, again, is orthogonal to the point about constrained vs unconstrained layouts (it makes no sense to constrain layout when plotting one network). And for the papers that had the choice between constrained and unconstrained layouts, your statement above is false:
(Rhemtulla, M., Fried, E. I., Aggen, S. H., Tuerlinckx, F., Kendler, K. S., & Borsboom, D. (2016). Network analysis of substance abuse and dependence symptoms. Drug and Alcohol Dependence, 161, 230–237. http://doi.org/10.1016/j.drugalcdep.2016.02.005)
Miri: “I’ve felt compelled to respond when our work is misrepresented and when my competence and integrity are publicly questioned.”
Are you talking about your paper that questions the competence and integrity of our two ex-graduate students who spent 4-5 years developing network models, vetting them carefully in numerous papers and simulation studies published in sophisticated technical journals like Psychometrica and Psychological Methods, and who developed software to enable researchers to use them? Because your paper clearly implies that these two graduate students—and their supervisors, colleagues, co-authors, and peers who supported them in these endeavours—have to be pretty big idiots. After all, they all missed a very basic insight that can be readily obtained by fitting network models to two datasets, and by looking at plots showing what variance decomposition is (cf. Forbes et al 2017): that network models are “plagued with substantial flaws”, “produce unreliable results”, and have “limited replicability” and “poor utility”. That can only imply a massive lack of competence on the side of these researchers. Or did you instead mean to imply a lack of integrity: that all these people know about these glaring issues, but ignore them because they are bad scientists?
Hi Miri,
1. You state that your “competence and integrity are publicly questioned”. I’m sorry but this statement is just ridiculous. Scientific criticism is NOT the same as questioning someone’s competence and integrity. You cannot expect to publish a paper that makes extremely strong (and often unfounded) claims and expect the people who’s work you criticise to keep their mouths shut. I get that you try to sell your paper by making strong claims. This is just how it works, right? But please stop complaining when someone calls you out for this.
2. You state that “the F-R algorithm is largely irrelevant to the key issues in the papers”. I’m not sure that I agree, but if you think that this is the case then it is even more frustrating that you cannot admit that you made a poor choice by using this algorithm. Also defending your choice by basically saying ‘everyone else does it” is a very weak argument.
3. You state that “This discussion is frustratingly unproductive”. Maybe if you don’t repeat your arguments over and over again the discussion would go somewhere.
4. You frequently refer to the “peer-reviewed exchange ” and by this hold up the peer review as a kind of shield of quality. For every reader somewhat familiar with psychometric network modelling it should be obvious that the quality control in this “peer-reviewed exchange” failed horribly, which is not that surprising, because you didn’t publish your methodological critique in a methodological journal.
Jonas
I also find this a “frustratingly unproductive” discussion and will try to opt out after this, as this discussion is going completely nowhere. Miri seems fixed on defending every little aspect of the paper(s) of interest, as well as to portray me or my colleagues as bullies. We simply show the fact that F-R is poorly suited for comparing networks and that showing the data in other ways, such as constraining the layouts, or plotting the data in heatmaps shows that the networks are much more similar as represented in the paper:
And for that, we are blamed for misrepresenting things and are called out for making outrageous and ridiculous points. As I also described in post-publication review on pubpeer, the original study simply does not show any “evidence that psychopathology symptom networks have limited replicability.” That is not a personal attack aimed at questioning your competence Miri, that is simply what follows from the data. You keep referring to your rebuttal, but it completely ignores several important points, as I already discussed in my pubpeer response:
In addition, the rebuttal only glosses over the high correlation between edge-weight parameters (again copied from my pubpeer response):
There, you only respond by saying that the points are discussed in the rebuttal and that in addition now your paper also shows evidence that permutation testing must be wrong.
The original paper simply does not show any evidence that network models are “plagued with substantial flaws”. At best, it shows that under LASSO weak edges (including the bridge symptoms) are somewhat unstable (which has been known for a long time), and that the full rank-order of centrality indices has limited replicability (which makes perfect sense, nobody would expect the rank order of factor loadings to replicate too). The only contribution in this entire exchange that I would fine, to borrow words from the other commentary, potentially interesting, is the PTSD literature review. But that is another topic entirely. To end, I would like to re-state places were we may find a middle-ground (text form with clickable links is in my pubpeer response):
I’ve been reluctant to join in on these exchanges; I’ve not been convinced of the value of social media for resolving nuanced issues.
However, I’d like to make an observation and take Sasha’s up on his points of agreements.
Based on what I’ve read here and elsewhere, my impression is that emotions are running high and that makes it difficult to have a scholarly debate. I don’t mean to point fingers. Depending on what perspective you bring to the table, what one paper says or what someone writes on a blog post may seem more or less reasonable or more or less outrageous. What I mean is, when emotions are high, it is hard to interpret what others have written charitably and take the others’ perspective. When that’s the case, misinterpretations and misattributions are likely, and it is too easy to focus on technicalities, thus missing the point (e.g., pointing out that the constrained plots from Rhemtulla, Fried, et al. [2016] used here to support one point are based on zero-imputation would only inflame that issue, and lead to more accusations and ill feeling). There’s too much latitude in this discussion. This could go on forever, and I think it would advantageous if it were to conclude before it goes further.
So, I will resist the temptation to try to argue, score points, clarify, or otherwise try to convince anyone to interpret our words in the way they were intended, and instead I will agree with the points Sasha outlines as potential middle ground. I’m glad to see that there is agreement on these points (as well as some others peppered throughout the exchanges), and I hope they get more widely disseminated and other users of these methods come to appreciate them as well.
I will not respond further here.
As much as I don’t want to come over rectifying mistakes on blog comments… My name is Sacha with a C, not an S 😉
Thanks for chiming in Aidan, and good to see there is common ground here. I forgot to mention the zero-imputation of skip structure. That, too, is an important problem that is now highlighted thanks to your paper, which I think marks another contribution to the field. Zero-imputation and improper handling of it is a huge problem in multivariate statistics we cannot ignore: logistic regressions, log-linear modeling (both of which Ising model is a special case), and tetrachoric correlations simply cannot handle such data in which marginal crosstables feature zeroes. That is evident from the extremely high correlations in the association networks (>0.9) and the unstable edges linked to the skip-items in bootstrap. Not really reported anywhere is that I also failed to estimate non-regularized Ising models (using IsingSampler) and that it is very hard to estimate factor models on these data. Finally, the skip-structure, true induced causal effects in the data, naturally bias all results: all networks simply show the skip-structure with some background noise. To this end, for all we wrote about this, we have yet to actually see a proper network computed on the NCS-R dataset that shows some interesting relationships rather than just the skip-structure (my hypothesis is that with proper handling of the skip-structure the replication between the datasets would be slightly worse as is now btw, as I judge the replication almost too good to be true now). I have been trying to get FIML tetrachoric correlations in OpenMx to work on this, but haven’t had much success yet. Perhaps we should just try Bayes. Would love to hear some thoughts on how to handle zero-imputed skip structures (let’s say in computing correlations alone, not necessarily networks)!
Thanks for chiming in Aidan, and I’m happy to hear you see some common ground (and was sorry to not run into you at ABCT last week). Skip imputation in the Rhemtulla et al. paper is a big issue, and I’m glad you brought that up. So let’s compare these two papers briefly, regarding (1) conclusions and (2) limitations.
1) Conclusions
All substantive findings from our paper should be interpreted with great care. As you surely have noticed, we do not draw strong conclusions in the paper the way Forbes et al. do. Main conclusions from the abstract of our paper: “When analyzed as networks, abuse and dependence symptoms do not function equivalently across illicit substance classes. These findings suggest the value of analyzing individual symptoms and their associations to gain new insight into the mechanisms of SUD”. Pretty carefully phrased.
In contrast, your paper draws very strong conclusions, which I have quoted above numerous times: “Popular network analysis methods produce unreliable results”, “Psychopathology networks have limited replicability”, “poor utility”, and, later, “current psychopathology network methodologies are plagued with substantial flaws”.
2) Limitations
In our paper, we clearly mention skip imputation as a limitation in the limitations section. Forbes et al., in contrast, do not acknowledge skip imputation as a limitation.
These seem two very important differences when comparing the papers. Now, I can only speak for myself here, but I readily admit that in early 2015 when writing this up, I had not been aware of the detailed issues skip imputation would pose to models based on correlations among items. I learned quite a bit about that in the 2 years since. But we do mention the issue as a limitation, despite it being an earlier paper than Forbes et al., and we draw weak exploratory inference from our analyses. In contrast, Forbes et al. do not mention nor acknowledge skip imputation as a limitation (not even in the rebuttal), nor do you draw weak inference from your analyses.
Pingback: 6 new papers on network replicability | Psych Networks
Pingback: 7 new papers on network replicability | Psych Networks
Pingback: Network stability part II: Why is my network unstable? | Psych Networks
Pingback: Estimating psychological networks via Information Filtering Networks | Psych Networks
Pingback: How to interpret centrality values in network structures (not) | Psych Networks
Pingback: Fixed-margin sampling & networks: New commentary on network replicability | Psych Networks