After Angelique Cramer and colleagues published the first paper on empirical networks for psychopathology in 2010, a few years of great enthusiasm followed, and psychological network science felt a bit like the Wild West: Everything was possible, and it was difficult to find qualified reviewers due to the novelty of the methods.

Recent years have seen more attention to simulation studies, methodological rigor, and a focus on stability and replicability of network models. Consistent with this trend, it is not surprising that two papers are in print at the same time now that focus on challenges of the network approach. These papers were written independently of each other, and while Sinan was kind enough to refer to our preprint, we were unable to cite their paper because we were not aware of it.
Challenges to the network approach: pitfalls, promises, new directions
The first paper, “Application of network methods for understanding mental disorders: pitfalls and promise”, was published by Guloksuz, Pries and van Os (from now on GPvO) in Psychological Medicine. The fairly concise paper is focused on four challenges: (1) a reductionist understanding of medicine and psychiatry, (2) a shortsighted view of signs and symptoms, (3) overlooking the limitations of available datasets, (4) and over-interpreting evidence (PDF).
The second paper, “Moving forward: challenges and directions for psychopathological network theory and methodology”, authored by me and Angelique Cramer1, is in press in Perspectives on Psychological Science. The challenges we discuss are: (1) The validity of the network approach beyond some commonly investigated disorders, (2) the definition of psychopathological systems and their constituent elements, (3) how can we gain a better understanding of the causal nature and real-life underpinnings of associations among symptoms, (4) heterogeneity of samples studied with network analytic models, and (5) a lurking replicability crisis in this strongly data-driven and exploratory field. Reviewers requested fairly extensive additions, so the paper ended up also including an introduction to network theory, and a brief overview of network psychometrics (PDF).
Instead of discussing these papers separately, point by point, this blog post has the goal to highlight some of the key messages. This means that several challenges remain unaddressed, but I wanted to give an overview rather than reiterate all individuals points. The blog post follows this structure:
- Complex dynamical systems, common causes, & oversimplifications
- A multilayer perspective on psychopathology
- Hybrid models: the complex reality of psychopathology
- The heterogeneity and limited validity of DSM categories
- Heterogeneity of clinical populations
- It’s complicated!
- Limitations of the DSM, and a call for transdiagnostic work
- What to include in networks? Sign & symptom limitations
- Biology!
- Symptom semantics
- What variables to include in network models
- Overinterpretation of evidence: networks as a cautionary tale
- Stability and generalizability of networks
- Causality and cross-sectional data
- The good old within vs between debate
- Ways forward
1. Complex dynamical systems, common causes, & oversimplifications
Psychopathology network theory — symptoms are correlated because they interact with each other causally — has been pitted against the common cause model in many prior papers. Examples often raised are depression on the one hand (insomnia → fatigue → concentration problems), and measles on the other where symptoms are passive consequences of a common cause.
We use an adapted version of this figure in our paper to explain this difference.
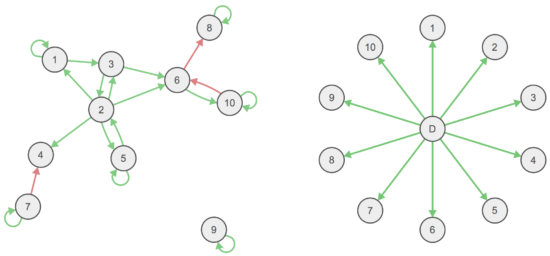
Both challenges papers take issue with this oversimplification, for the following reasons.
1.1 A multilayer perspective on psychopathology
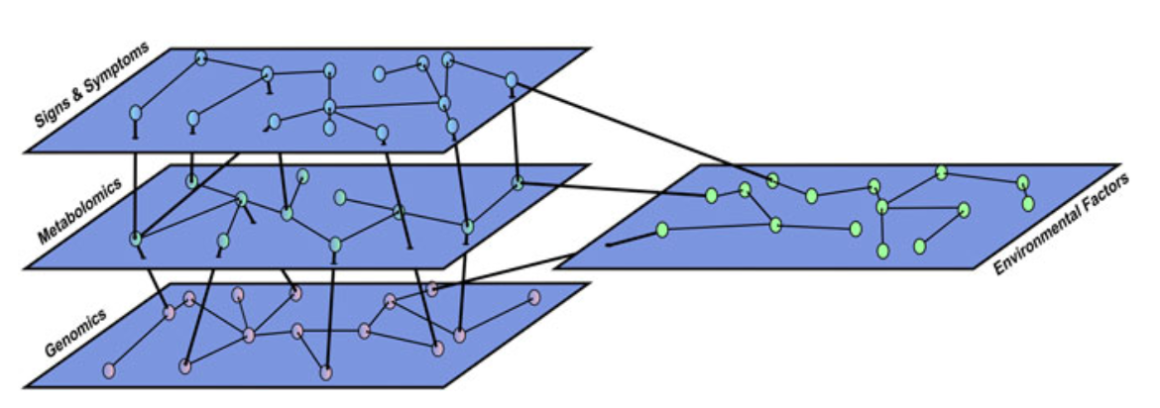
GPvO argue that the separation between conceptual network and common cause models is somewhat artificial, and re-introduces a sort of Cartesian Dualism (mind vs matter), which is not helpful in advancing clinical science. Instead, they propose a multilayer network to explain the intricate interactions among more biological and more psychological variables (I say ‘more’ here to avoid reintroducing this dualism in the explanation of GPvO’s work):
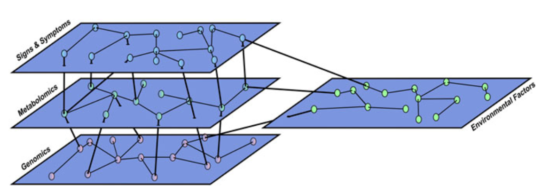
From this perspective, so GPvO, lung cancer and major depression might be very similar, the only difference being “that we have a deeper understanding of underlying biological abnormalities in the former […] while psychiatric classification is stuck at the level of signs and symptoms.”
In my mind, this is an interesting idea and ties into similar work of interactions between layers of pure biology all the way to pure psychology, with plenty of intermediate layers. Causality among layers in this sort of model is usually very hard to disentangle because it can go both ways (cf. epigenetics).
1.2 Hybrid models: the complex reality of psychopathology
Our paper highlights a different shortcoming: that the distinction between pure network and pure common cause models oversimplifies the complex reality of mental disorders. Is it really likely that there are no common causes for symptoms at all? And, on the other hand, do residual correlations in factor models not show us that the common cause model might better be complemented by a network model that can causally explain these associations?
We propose hybrid models in which both common causes and networks work together. For instance, a traumatic experience could explain the initial onset of PTSD symptoms, while a network takes over in the maintenance phase of the disorder. We discuss this for other disorders such as depression as well, and go through a few more complex examples, including e.g. moderators.
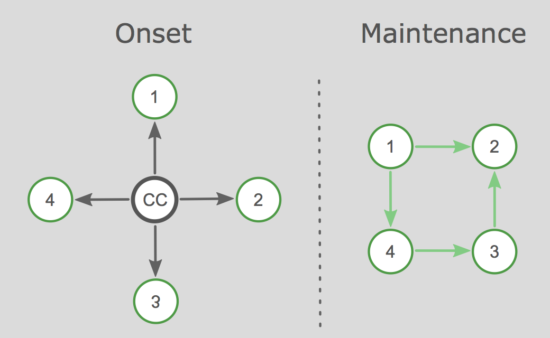
2. The heterogeneity and limited validity of DSM categories
2.1 Heterogeneity of clinical populations
In both papers, the heterogeneity of mental disorders features prominently, which can be difficult to capture in network models. We discuss this in detail, advance network mixture models as a possible solution to heterogeneity in cross-sectional data, and discuss how recent advances in time-series models can be used to tackle the issue of heterogeneity in experience sampling data with several timepoints a day collected over several weeks. In such data, multilevel models allow for e.g. 3 patients to have different individual networks, and group-level networks and variability networks can be used to give complimentary information on how similar these networks are:
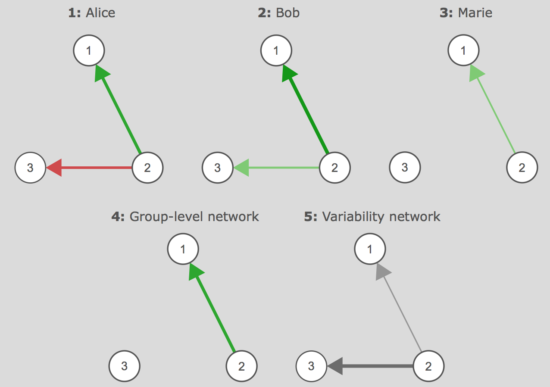
We also discuss other possibilities, such as clustering people in time-series networks, or modeling approaches such as GIMME developed by Gates & Molenaar that may offer insights into heterogeneity.
2.2 It’s complicated!
Heterogeneity connects to another point: while statistical models in cross-sectional data – including network models – are capable of investigating hypotheses at the group level (e.g., women have more strongly connected depression networks than men), such results may not generalize to all individuals the population consists of, the same way the group-level information “men are taller than women” does likely not hold for all individual people (my girlfriend, for instance, is 187cm tall).
We bridge the gap between (a) network models, (b) common cause models and (c) hybrid models on the one hand and nomothetic vs idiographic analyses on the other by concluding:
“Therefore, an equally interesting and possibly more complicated question is: Which of the three models described above fits the psychopathology of a given person best? MD, for instance, could stem from a common cause (e.g., brain pathology), a network model (e.g., vicious circles between negative thoughts and emotions; Beck et al., 1979), or a hybrid model (e.g., a network following severe adversity), depending on the specific individual and her or his specific circumstances […]. This view stresses an idiographic perspective on mental health research and acknowledges that only embracing the heterogeneity of diagnostic categories will enable us to make true progress toward personalized medicine (Kramer et al., 2014; Molenaar, 2004).”
2.3 Limitations of the DSM, and a call for transdiagnostic work
GPvO also stress the topic of heterogeneity, and add that current datasets are often limited by design. For instance, while sad mood and anhedonia often come up as the most central symptoms in depression network studies, the reason could simply be that this is because they are necessary for a diagnosis, and because people without these symptoms are not included in clinical studies based on diagnostic interviews.
Both teams make the point that DSM categories have received considerable criticism in recent years. Current diagnoses may be somewhat reasonable clinical descriptions and a judicious starting points for network studies, but network theory will not be able to make up on its revolutionary promises if the datasets analyzed are not transdiagnostic in nature. One of the most central tenets of network theory is that “problems attract problems”, and this holds across diagnostic boundaries, and offers a different explanation for comorbidity than the notion that e.g. major depression and generalized anxiety disorder are the result of two distinct etiologies.
As GPvO put it:
“However, it is difficult to understand the logic of inhibiting the potential of datasets by deliberately confining the network to boundaries of DSM […]. It is reassuring to see that this common practice has been changing gradually in more recent studies. Otherwise, one might ask that if network theory has no other option but to play the game with the rules of DSM, how will it turn out to be the promised game-changer?”
3. What to include in networks? Sign & symptom limitations
Many studies in this field have re-analyzed existing datasets, which are largely based on rating scales or DSM diagnostic criteria. This is, as GPvO say above, a limitation by design. Many other variables might be interesting inclusions in network models, including environmental and biological variables.
3.1 Biology!
GPvO write a general critique of the usefulness of signs and symptoms, arguing that they are “usually non-specific, commonly subjective, qualitative or difficult to quantify, and therefore, inadequate to unequivocally diagnose a patient”, and that clinical classification is therefore inherently limited. They make the case that insights into the underlying pathoetiology will lead to better classification. I would add here that many symptoms are featured multiple times in the DSM (fatigue and insomnia numerous times), and also shared with medical diagnoses, which makes them fairly bad candidates for any “classification” (with the idea that symptoms indicate an underlying disorder).
However, I am not sure this is a limitation of network theory, which in its purest form posits that a disorder such as major depression is nothing but an emergent property of symptom interactions. I believe this position to be overstated, and we talk about this in great detail in the hybrid model section of our paper and discuss that many local common causes exist that make symptom endorsement more likely, and speculate how this could work conceptually and statistically. Nonetheless, from this perspective, there would be no “underlying pathoetiology” as suggested by GPvO — the notion of an underlying disease is based on the common cause notion that a disorder is causally responsible for symptom covariation, in which case networks are not the models you want to use to look at your data. Such a “purist” network perspective would not rule out biological changes in patients with mental disorders of course, merely posit that these are not causal for symptomatology. GPvO offer a different approach, however, by suggesting a multilayer network with causal relationships between e.g. a ‘biological’ network and a ‘psychological’ network. Causes seem to go both ways, since GPvO draw undirected edges, which strikes me as very plausible. In this case, a psychological network can have an underlying biological network, although underlying here has different causal implications. GPvO also stress that biological and psychological variables may best be modeled together, and specifically mention inflammation and depression symptoms 2.
3.2 Symptom semantics
In our paper, we first discuss the term ‘symptom’ more generally, showing that it actually makes little sense in the context of networks, given common dictionary definitions. The Cambridge dictionary, for instance, states:

For this reason, other researchers like Donald Robinaugh and Richard McNally have suggested to instead use the term “elements”. Instead of focusing on underlying biology like GPvO, we instead argue that many other psychological variables can be included in network studies, and that “the traditional conceptualization of the relation between mental disorders and symptoms has granted symptom variables a certain importance above and beyond other clinical variables”.
3.3 What variables to include in network models
We move on to define dynamical systems, and discuss important additions for future studies (both variables in the system and variables in the external field). The transdiagnostic arguments provided above of course speak for including symptoms of numerous disorders, and beyond symptoms, important variables could be impairment of functioning, cognitive processes (e.g. self-esteem or a sense of self-efficacy), distress, approach or avoidance behaviors, positive or negative social interactions per day, rejection events, physical activities, and substance abuse.
Further, we discuss the consequences of failing to incorporate relevant variables in dynamical systems, and if items should be combined or not in case they are somewhat similarly phrased. In the first case, spurious edges among the other items in the network will likely emerge, and we require both statistical approaches and sound theory to define what we should model. In the second case, we describe the notion of topological overlap in more detail which might help to decide whether similar items (e.g. ‘feeling blue’ and ‘sad mood’ from the CES-D depression screener) should be featured in a network separately, or better combined to one single node:
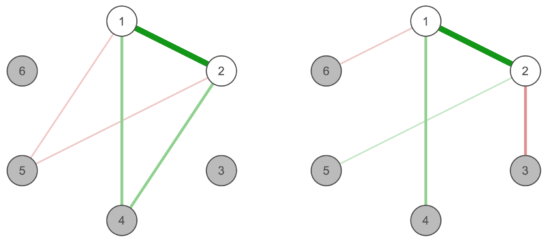
In the left case, the two white items are highly correlated and occupy the same “position” in the network, which would suggest they ought to be merged. In the right case, they show differential associations (akin to weight and height that would be highly correlated, but are different things and would show different edges with variables such as “do I want to lose weight” or “can I see far in concerts”).
GPvO articulate a somewhat related challenge by criticizing that network researchers often combine items such as weight gain and weight loss, or psychomotor retardation and agitation, into one aggregate score, following the logic of the DSM. But this makes little sense, and research sides with the position of GPvO here3.
4. Overinterpretation of evidence: networks as a cautionary tale
Networks provide us with beautiful visualizations of models and data, and as my colleagues in Groningen say: there is a certain ‘Rohrschach’ element to looking at them. This is true, and more caution is required when interpreting networks (I have also devoted a recent blog post to this topic). There are several issues here.
4.1 Stability and generalizability of networks
An important topic we have worked on in the last years is network stability and generalizability, and Angelique and me also discuss it in-depth as core challenge in our paper. We talk about overfitting and exploratory models, and make the point that parameter (or model) stability is necessary, but of course not sufficient for replication of models across datasets. This means that we should put focused effort into testing the stability of our network models (and, of course, all other statistical models such as factor models), and in a second step investigate whether network models replicate across datasets or not.
How exactly is this related to visualizations of data? In our paper, we provide a toy example where we call for readers to write a brief network paper with us.
“So let us write a quick paper together to see why stability matters. We estimate a network of 17 PTSD symptoms in a sample of 180 women with posttraumatic stress disorder: A strong edge emerges between 3 and 4, representing a clinically plausible association between being startled easily and being overly alert. We also observe a negative edge between symptoms 10 and 12 and conclude that people who do not remember the trauma are less likely to have trouble sleeping (and vice versa). In a second step, we investigate the centrality (connectedness) of nodes (Opsahl, Agneessens, & Skvoretz, 2010). In our example network, node 17 has the highest degree of centrality (1.25) and node 7 the lowest (0.65). We now finalize the paper and suggest that future studies should pay specific attention to edges 3–4 and 10–12 and that targeted treatment of node 3 may achieve the greatest benefits for patients. Success!”
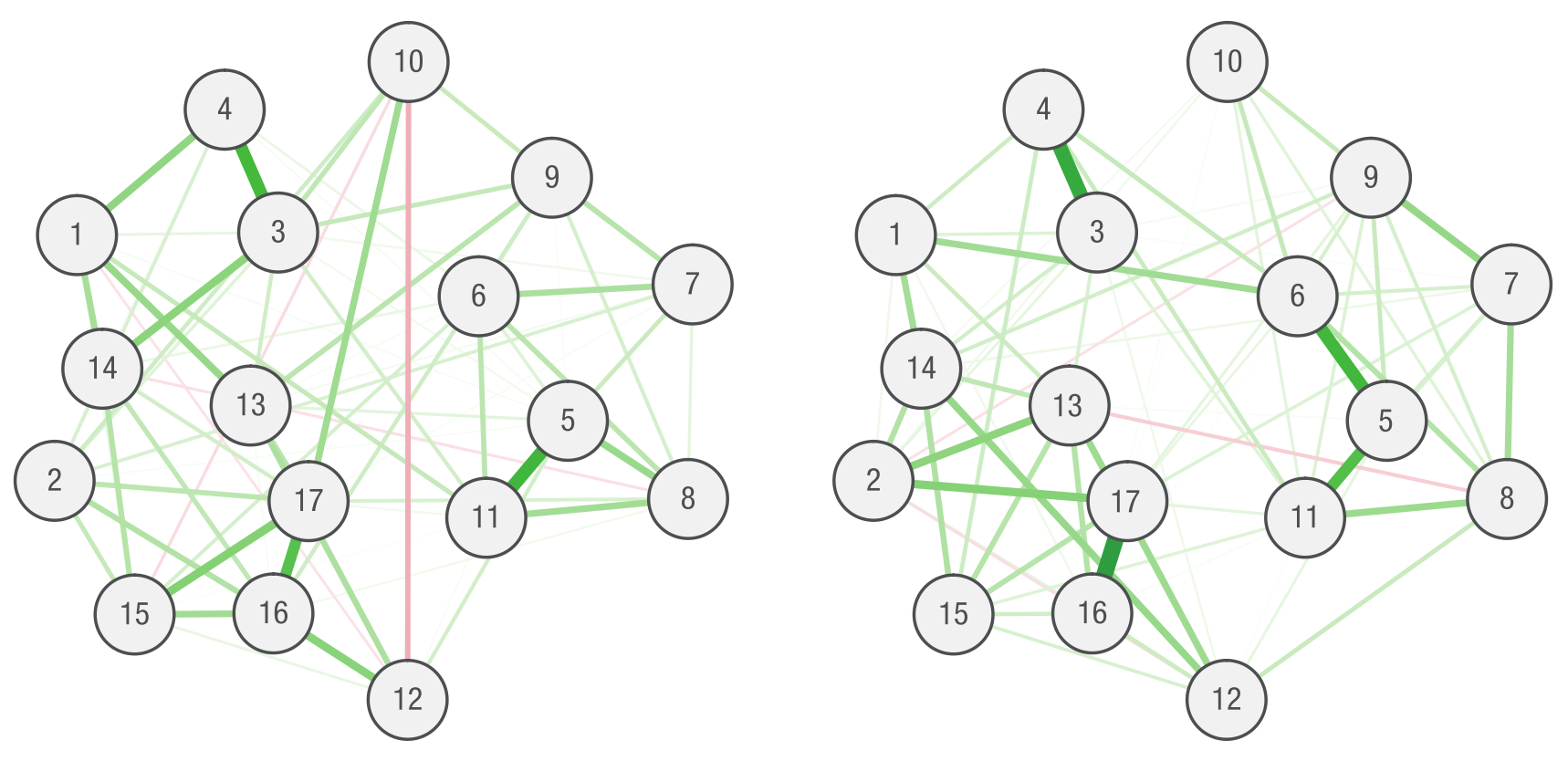
We then “stumble” across a second dataset that is similar to the first. The network in this second dataset is quite different, the reason being that parameters are not estimated accurately in either due to the low sample size. In essence, a lack of stability also means a lack of replicability, and we show that paying attention to parameter stability is a safeguard against drawing wrong conclusions that the data do not support.
4.2 Causality and cross-sectional data
Both papers stress the importance of understanding the limitations of data. Cross-sectional data are cross-sectional, and that is that. While there are different opinions on whether one can infer a causal skeleton from such data, e.g. via directed acyclical graphs, there is no question that one cannot infer causality from such data. And while we are at it, you can neither infer causality from time-series data (ask Granger). While I tend to disagree with the stricter critics of the network approach that the literature drowns in papers of causal inference in cross-sectional data, it is true that research has largely focused on cross-sectional data, which is somewhat of a mismatch with network theory that is inherently dynamic (i.e. temporal).
Our paper also features a chapter on how we need to move forward to truly advance networks as causal systems, with a focus on experimental manipulations and mechanistic explanations:
If insomnia → fatigue is the true model underlying the observed correlation between insomnia and fatigue, intervening on insomnia should reduce subsequent fatigue. In contrast, this intervention will not be successful if a common cause underlies the two symptoms, in which case only intervening on the common cause will successfully cure both symptoms.
4.3 The good old within vs between debate
Both papers also highlight, in different levels of detail, that between-person and within-person analyses are different and answer different questions. Some of the reviewers of our paper argued strongly against between-person network research in general, but we do not agree here, a point we make repeatedly in the paper. Both views are interesting and valid, but we should only draw conclusions that can be supported by the data.
5. Ways forward
In our paper, we discuss possibilities to develop network theory for other mental disorders that have not been investigated so far, and end with a call for action for both methodologists and researchers. Methodologists can advance the field by looking into (1) confirmatory network modeling, (2) network mixture models, (3) the statistical comparison of latent variable models to network models, (4) network a priori power analysis 4, and (5) more non-technical tutorial papers. Clinical researchers can look into (1) development of network theory for different disorders, (2) collecting data that can be readily analyzed using network approaches (we don’t quite think about this enough before collecting data), (3) and always testing and reporting stability and accuracy which is necessary for drawing inferences about replicability.
GPvO end with a call for (1) more transdiagnostic research, (2) proper replication studies, (3) more cautionary interpretation of network studies in general, and (4) focused assessment of experience sampling methodology to collect time-series data.
Here comes a last final concluding sentence, I guess. But since I don’t expect anyone to read this far, I will just skip it 😉5.
References
» Guloksuz, S., Pries, L.-K., & Van Os, J. (in print). Application of network methods for understanding mental disorders: pitfalls and promise. Psychological Medicine, 1–10. 10.1017/S0033291717001350. (PDF)
» Fried, E. I., & Cramer, A. O. J. (in print). Moving Forward: Challenges and Directions for Psychopathological Network Theory and Methodology. Perspectives on Psychological Science. 1-22. DOI 10.1177/174569161770589. (PDF)
- In German we say “The donkey refers to himself first” for such a sentence, but here this is just author order, not me being incredibly impolite ;).

- This is great news, seeing that a student of mine has been working on exactly this project for over half a year, and we’re ready to submit the paper soon!
- I have actually empirically investigated this in several papers, and combining depression symptoms opposites is not justified due to their differential associations with numerous outcome variables such as impairment of functioning (paper1, paper2, paper3).
- Sacha Epskamp tackled this challenge of a priori power analysis in our tutorial paper on network stability and accuracy after Angelique’s and my challenges paper was accepted at Perspectives, so unfortunately we could not discuss this in the challenges paper.
- And for the one person who actually read all of this: talk to me at the next conference, I’ll buy you an icecream!
Hi Eiko,
I greatly benefitted from the insights of the networks approach ever since I learned about it around a year ago and it complements related interests I have in various fields. However, I have two basic questions that have been bothering me for a while now and so far haven’t been able to find good answers/solutions. Because this post is about pitfalls, challenges, and future directions, I wanted to bring them up here:
1) Why partial correlation? I know what they are and how attractive it is to ‘isolate’ things. But what’s actually wrong with using straightforward Pearson correlations to construct weigthed networks?
2) Heterogeneity/Reliability/Sample size? I get the issue, re-read relevant sections in your article Fried & Cramer, and I follow related discussions in neuroimaging (part of my work is in that field). Specifically, in functional brain connectivity networks are constructed from bascially correlations (or other measures – including partial corrs) between different regional time series. At the individual level, this also is affected by lots of noise. However, the network here is estimated for each individual and can then be aggregated across people, which will help to ‘beat down the noise’. What I don’t quite understand is where exactly a different issue arises with psychological networks (or is neuroimaging just blind to it? [rhetorical q – there are many papers on reliability coming out over the past years and recently particularly for connectivity]). Isn’t it possible to perform significance testing with multiple comparison correction for each edge? Or what about a null-model based on data from normals (whatever that means) and compare the disordered against? I must be missing sth. … As said, I totally get the importance of good measures (minimize noise to see clearly) and the problem of heterogeneity (even noise-free measurement will still give blurred solutions if people differ), but why is this soo special to networks? Your writing seems to imply that’s because so many parameters have to be estimated? But why do they have to be estimated instead of be ‘measured’ (see q. above)?
Thanks a ton – awesome work
Ralf
Hi Ralf!
1) If you want to predict mortality at time 2 by depression at time 1, you arguably want to control for a few things, such as gender, age, and maybe comorbid conditions and health behaviors such as smoking. Visualizing the correlation table as a network here — which is not a model of any kind — will not get you anywhere; you would likely find that all variables relate to mortality. But a multiple regression that is able to account for shared variance among the predictors might give you the answer that depression does not predict mortality once you end up controlling for other variables (this is actually what we found in a recent study). This is one of the reasons we use regularized partial correlation networks. We are interested in conditional (in-)dependence relations among items. If the true model among 3 items is A → B → C, and you have cross-sectional data, your bivariate correlations will reveal correlations between all variables. Only the conditional dependence, however, will truthfull return A — B — C without a connection between A and C.
Verena Schmittmann et al. have a kickass paper in Plos One showing why neuroimaging should move to partial correlation networks.
For more information on this topic, see work by Pearl (e.g his 2000 or 2001 book on causality), and the two papers on the Ising Model and the Gaussian Graphical Model, that also explain the difference between partial correlations and regularized partial correlations in more detail.
2) I’m not quite sure I get the second question, and I admit I don’t know how time-series networks look in brain connectivity research. But in principle, the huge difference between e.g. social networks and psychological networks is that in social networks, both nodes (people) and edges (e.g. friendships) are observed, while the latter need to be estimated in our case (and I assume in your case, too). If you have 30 people, 10 measurement per day for 2 weeks, and 10 items of interest, you can calculate the crazy amount of associations you need to estimate, especially if you want to get both contemporaneous and temporal networks out of the data (see reference manual; note also that people and timepoints give you power, while more items reduce your power). Maybe in brain networks you just have a ridiculously high number of timepoints, which gives you many orders of magnitude more power?
The same parameter issueholds for cross-sectional data: there is no observation of edges, we need psychometric models to estimate these, and that is what regularized partial correlation networks do. If you have 30 items, and you are interested in conditional dependence relationships, you need 30*29/2 edge parameters (because you regress every node on every other node; so for 3 items A B C you estimate A~B+C; B~A+C; C~A+B; then choose a specific rule how to deal with the fact that A~B and B~A can be estimated differently; and then you draw the graph). Hope this makes sense, please let me know if this remains unclear.
HI Eiko,
thanks. Very helpful – though I’ll need a bit time to digest. Just a brief follow-up.
Let’s start with the 2nd question, which indeed I am myself was not fully clear how to ask: I see how estimating many noise-laden things will bring you in trouble. Agreed. You mention that the huge differences between social networks is that edges are observed (friendships), whereas you have to estimate them. Okay, if you use the full partial-correlation permutation (A~B+C etc) then you indeed have a lot of estimation to do. But what if you take the raw correlations. Aren’t they somehow ‘observed’ (measured as the strength of association between eg. sleep issues and lack of concentration – similar to asking A how much friend s/he is with B and vice versa [1-7]?)? Or would you say there is still a difference between a low-noise observation and a more noisy measurement (estimation?) of the association between two fMRI timeseries?
I guess that brings us back to the question #1, i.e. whether you want/should start getting into (regularized) partial correlations. Thanks for the Schmittmann paper, which I had not known and indeed liked. My reservation, however,is that you can find about as many papers arguing the opposite and backing that up with simulation and limited empirical data (e.g. Kim, Wozniak, Mueller, Pan, 2015; Zalesky, Fornito, & Bullmore, 2012; or Smith, Miller, Salimi et al., 2011). Okay, that is not in itself a great argument, but it casts at least a little doubt on the point that partial correlations are the demonstrated go-to method. I can see how in your example about mortalityT2 ~ depressionT1 + age + gender you want to control for these things. However, in the more exploratory search of how different symptoms hang together as a network, doesn’t this kind of model come with assumptions? In particular, it seems to ignore interactions or higher-order relationships (beyond pairs) that seem theoretically plausible to me, especially from a dynamical systems perspective. E.g. this article blew my mind (https://arxiv.org/pdf/1608.03520.pdf) when I read it. Again, I am not so confident in my knowledge to make any strong statement here, but the bottom line to me seems to be: (regularized) partial correlations are just an analysis method. They don’t introduce new information to the data. Any added insight comes from either the fact that their assumptions and the transformations they introduce are plausible or that they let you observe the data from a perspective that is otherwise obstructed (as in your A-B-C example). It seems to boil down to that ‘conditional independence’ issue which I will need to think about more. However, given the typically larger complexity of real-world data, my intuition has always been to stay close to the phenomenon and I didn’t yet fully grasp why partial corrs are necessary. I agree that in your mortality prediction and A-B-C examples that isolating/controlling are desirable, but in case of uncovering the network structure of “depression” symptoms in the first place, I was wondering about whether all the challenges you mention and about potential downsides of an exclusive focus on partial correlation.
Thanks again for the reply. Your papers have tremendously enriched my thinking
Best r
Thanks for following up Ralf!
“Okay, if you use the full partial-correlation permutation (A~B+C etc) then you indeed have a lot of estimation to do. But what if you take the raw correlations. Aren’t they somehow ‘observed’ (measured as the strength of association between eg. sleep issues and lack of concentration – similar to asking A how much friend s/he is with B and vice versa [1-7]?)? Or would you say there is still a difference between a low-noise observation and a more noisy measurement (estimation?) of the association between two fMRI timeseries?”
—— I am conceptually not interested in correlations — I do not want a fully connected network where every symptom at time t predicts time t+1, I want the information which symptoms at t predict t+1 symptoms controlling for all other associations. I am interested in unique variances, the same way I am interested in predicting mortality by all my relevant variables in one prediction model. Univariate prediction will give you is spurious relationships, many of which would disappear once controlling for other variables. Regarding the question if a correlation is observed: not in a statistical sense. Your dataframe has the item responses, and you can estimate a mean of an item, or a correlation, no? From how I’ve seen the words observed vs estimated seen, a correlation is estimated. Regarding fMRI timeseries, I’m afraid I can’t answer that question, sorry. I really don’t know what models you are using in that field for time series estimation.
Regarding your second point:
“My reservation, however,is that you can find about as many papers arguing the opposite and backing that up with simulation and limited empirical data”
—— Cool, will have to check this out. The point we are making is conceptual, not statistical (nothing wrong with correlations, but they don’t give us what we’re looking for, based on the assumption where our data come from).
“However, in the more exploratory search of how different symptoms hang together as a network, doesn’t this kind of model come with assumptions? In particular, it seems to ignore interactions or higher-order relationships (beyond pairs) that seem theoretically plausible to me, especially from a dynamical systems perspective. E.g. this article blew my mind (https://arxiv.org/pdf/1608.03520.pdf) when I read it.”
—— Absolutely, this is a big limitation of pairwise estimation (i.e. both correlation and partial correlation): we miss higher order interactions. A PhD student in our lab is looking at interactions at the moment, and it looks like there is a considerable amount of higher order stuff happening in the data.
“(regularized) partial correlations are just an analysis method. They don’t introduce new information to the data.”
—— If your data come from a unidimensional factor model — if “g” really causes all subtests of intelligence — then a factor model is the appropriate model to fit to your data. I’m not sure it “introduces new information”, but factor loadings are actual estimates and reflect the data, while other models would not adequately reflect the data.
“I was wondering about whether all the challenges you mention and about potential downsides of an exclusive focus on partial correlation.”
—— I wish we would have discussed this more in the paper, you’re absolutely right that this deserves more thought.
Another point I just discussed with Sacha Epskamp: if your data come from A→B→C, your partial correlation network will recover this structure A—B—C, but your correlation networks will be fully connected. If your data come from a latent variable model, your partial correlation structure and your correlation structure will be fully connected. That means that the partial correlation network is different from different data generating mechanisms (at least in many cases), while the correlation network simply doesn’t allow for any of such inferences.
Pingback: A summary of my academic year 2017 – Eiko Fried
Pingback: How to interpret centrality values in network structures (not) | Psych Networks
Pingback: Fixed-margin sampling & networks: New commentary on network replicability | Psych Networks